Ultrasound and Serology For Diagnosis of Human Cystic Echinococcosis in An Endemic Zone in China
Article Information
Jian Liu1,2, Qi Gao3, Dan Li1,2, Yu Feng4, Wenhua Ning5, Yanqing Dong3, Jingyi Li1,2, Wenjun Xu6, Deli Xin1,2*
1Beijing Tropical Medicine Research Institute, Beijing Friendship Hospital, Capital Medical University, Beijing 100050, China
2Beijing Key Laboratory for Research on Prevention and Treatment of Tropical Diseases, Capital Medical University, Beijing 100050, China
3School of Public Health, Capital Medical University, Beijing, 100060, China
4Parasitology Institution, CDC of Gansu Province, Lanzhou, 730000, China
5CDC of Xiahe County, Xiahe, Gansu Province, 730000, China
6Beijing Electric Power Hospital,Capital Medicine University, Beijing,100073, China
*Corresponding Authors: Deli Xin, Beijing Tropical Medicine Research Institute, Beijing Friendship Hospital, Capital Medical University, Beijing 100050, China and Beijing Key Laboratory for Research on Prevention and Treatment of Tropical Diseases, Capital Medical University, Beijing 100050, China
Received: 25 March 2019; Accepted: 16 May 2019; Published: 20 May 2019
Citation: Jian Liu, Qi Gao, Dan Li, Yu Feng, Wenhua Ning, Yanqing Dong, Jingyi Li, Wenjun Xu, Deli Xin. Ultrasound and Serology For Diagnosis of Human Cystic Echinococcosis in An Endemic Zone in China. Archives of Microbiology & Immunology 3 (2019): 050-061.
Share at FacebookAbstract
Cystic echinococcosis (CE) is a public health concern in China. However, the incidence and prevalence of CE in the Chinese province of Gansu is not well known. CE diagnosis is often limited to ultrasonography (US) and serological tests. We have assessed the use of these two approaches for diagnosis of CE in the Xiahe county (Gannan prefecture, Gansu province), an endemic area in the northwest of China. This was a community-based cross-sectional study involving 972 people selected by multi-stage cluster random sampling. CE was investigated by US and serology, using a commercially available ELISA test. Eighty-four individuals tested positive for serology, out of whom 74 (88.10%) were 21-60 years old. Twelve individuals were diagnosed by US. Overall, seroprevalence was 8.63%, and the rate of individuals diagnosed by US was 1.23%. Seroprevalence was higher among females than males (10.16% vs. 6.5%), while the rate of individuals diagnosed by US was slightly higher among males (1.22% vs. 0.89%). Our results demonstrate that CE remains prevalent in the Gansu province. Serological monitoring can be a valuable tool for detection of infected individuals.
Keywords
Ultrasound; Sera,; Screening; cystic echinococcosis; Gansu province; Endemic
Article Details
Introduction
Cow’s milk allergy (CMA) is a IgE-mediated immunologically adverse reactions, which affects about 2.5% of newborn infants in the first year of life [1-2]. Although double-blind, placebo-controlled food challenge(DBPCFC) is considered as gold standard for food allergy diagnosis, the application of DBPCFC in pediatric clinical practice is still limited due to its high-risk and time-consuming procedure [3-4]. Therefore, in-vitro measurement of cow’s milk-specific IgE(sIgE), has been widely used in clinical practice as an alternative option for allergen laboratory diagnosis, to provide information on food allergen sensitization [5-6].
The composition of cow’s milk is very complicated, there were much evidence to show being sensitized to some certain cow’s milk components has proven to be more prone to a poor outcome of CMA [7-10]. Casein, as the most abundant protein, has been proven to be both more antigenic and allergenic than other cow’s milk proteins [11]. Children with persistent CMA had an elevated levels of casein specific IgE antibodies (Cas-IgE) than those with transient CMA, and patients in this population were tend to have a higher risk of severe anaphylaxis and a delayed course of food reintroduce [12,13]. In addition, Machteld M et al. [14] study has proven distinctive properties of cow’s milk specific-T cells responses existing among patients with persistent cow’s milk, cow’s milk-tolerant and nonallergic control subject, which suggesting that the potentially critical role of cow’s milk allergen-specific T-cell-mediated reactivity of children with or without CMA. Accordingly, in order to clarify the detailed role of casein-specific T responses in cow’s milk allergy, this study was designed to compare the casein-specific T cells response among CMA children with or without casein sensitization, basing on expression of T cell-related transcription factor, antigen-specific proliferation, and the production of T cell related-cytokine.
Material and Methods
Subjects
A total of 25 children with IgE-dependent CMA were recruited from outpatient clinic in Beijing Children’s Hospital from January to August of 2017, including 14 CMA patient with a positive sensitization to casein component, 11 CMA patient being sensitized to cow’s milk, but a negative sIgE for casein component, and another 10 atopic control subjects without CMA were also included as negative control group in this study. The diagnosis of CMA was based on the basis of a compatible and convincing clinical history relation with immediate cutaneous reaction after intake of cow milk. All enrolled subjects were fully informed about the purpose of this study and nature of the study, which were approved by the medical ethnic committee of Beijing Children’s Hospital. Written informed consent was obtained from all parents/guardians of all the participants.
Determination of total and allergen-specific IgE antibodies
Serum levels of total IgE, specific IgE antibodies (sIgE) to cow’s milk(CM-), casein(Cas), α-lactalbumin(α-La) and β-lactoglobulin(β-Lg) were measured using ImmunoCAP® (Phadia AB, Uppsala, Sweden). Any sensitization was regarded as positive when specific serum IgE levels were noted as being greater than 0.35 kUA/L against the tested isolated cow’s milk protein. The comtmercially available tests and reagents were used according to the instructions from the manufacturer.
Peripheral blood mononuclear cells (PBMCs) and total CD4+ T lymphocytes isolation
PBMCs were isolated from heparinized venous blood by density-gradient separation (Ficoll-Paque PLUS; GE Healthcare, Barcelona, Spain). Blood was diluted 1:2 with phosphate buffered saline (HAOYANG, Tianjin, China), then centrifuged using Ficoll density gradient solution(HAOYANG, Tianjin, China) at 2000 rpm for 20 min at 4 ? to isolate the PBMCs. Resting CD4+ T cells were positively selected from PBMCs using magnetic CD4 MicroBeads (Miltenyi Biotec) or negatively selected using a human CD4+T cell enrichment kit (StemCell Technologies). The purity of CD4+T cells was confirmed by using the FACScan system (BD Biosciences).
Cell proliferation rate evaluation upon in-vitro allergen-specific stimulation
PBMCs (2×106 cells/mL) were cultured in vitro for 7 days at 37 °C in 5% CO2 with ?-MEM medium alone as negative controls, or with 200 μg/mL mixture of purified alpha-, beta-, and kappa-caseins (Sigma-Aldrich, USA), or with 4 μg/mL phytohemagglutinin (PHA) (Sigma-Aldrich, USA) was used as positive control. Cow’s milk allergen-specific T cells proliferation status was identified using ex vivo carboxyfluorescein succinimidyl ester (CFSE) dilution before antigenstimulation with modifications to the method described by Turcanu et al. [16]. Briefly, PBMC samples were labeled with 5 mmol/L CFSE for 10 min at 37 ?.
Cytokine profile analysis
A custom designed MILLIPLEX MAP kit (Millipore Corporation, Bilerica, MA, USA) and the Luminex xMAP 200 System (Luminex, Austin, Texas, USA) were used to quantify Th1(IL-12, IFN-γ), Th2(IL-4, IL-13), Th17(IL-17, IL-23) and Treg(IL-10, TGF-β) in patients’ serum and in cell culture supernatant following manufacturer’s instructions. Briefly, the samples were thawed and incubated with cytokine-specific antibody-coupled beads, followed by incubation with indicator antibodies. After incubation, samples were subjected to the Luminex 200 System. All standards and samples were run in duplicates. Mean Fluorescent Intensity (MFI) data was collected and analyzed using a weighted 5-parameter logistic or spline curve-fitting method for calculating cytokine/chemokine concentrations (pg/ml). Results are expressed as amount of each cytokine detected after the stimulation with casein minus the amount detected after stimulation with the negative control.
RNA preparation and quantitative reverse transcription-polymerase chain reaction
Total RNA from PBMCs or isolated CD4+ cells were extracted using RNeasy plus mini kit (Qiagen) according to manufacturer instruction. RNA template was quantified using a NanoDrop ND1000 instrument (Thermo Fisher Scientific). RNA was converted to cDNA using PrimeScriptTM RT reagent Kit (TAKARA, Dalian, China) according to the manufacture’s instruction. Quantitative reverse-transcription PCR (qPCR) was performed using a SYBR Premix Ex Taq kit (TaKaRa, Dalian, China), following the manufacturer’s instruction. Primer specific for tested gene were listed in Table 1. The amplification program used was: 1 cycle of 10 minutes at 95 °C, 40 cycles of 15 seconds at 95 °C, and finally 1 cycle of 1 minutes at 60°C. All reactions were performed in triplicate. The mean value of the replicates for each sample was expressed as the quantification cycle (Ct). The Ct data were normalized using GAPDH and the relative gene expression level were calculated with the 2??Ct method.
|
Gene name |
Forward primers (5’-3’) |
Reverse primers (5’-3’) |
||
|
T-bet |
GGGAAACTAAAGCTCACAAAC |
CTCTCCGTCGTTCACCTCAAC |
||
|
FOXP3 |
CAGCACATTCCCAGAGTTCCTC |
GCGTGTGAACCAGTGGTAGATC |
||
|
ROR?t |
CTCAAAGCAGGAGCAATGGAA- |
AGGGAGTGAGAAGTCAAAGA |
||
|
GATA3 |
CGGCTTCGGATGCAAGTC |
GTCGAGGTTGCCCCACAG |
||
|
GAPDH |
AGCCGAGCCACATCGCT |
CAGCCCTGGTGACCAGGC |
||
Table 1: Primers used for real-time quantitative PCR
Statistical analysis
Statistical analysis was performed using SPSS version 19.0 statistical software (IBM, Armonk, NY, USA) and graphs were generated using the prism software (GraphPad, La Jolla, CA, USA). Qualitative variables were compared between the groups by chi-square tests and continuous variables were analyzed by the nonparametric Mann-Whitney U test for unpaired comparisons. P values of less than 0.05 were considered significant.
Results
Study subjects
A total of 25 children with IgE-dependent CMA were recruited in the present study, including 11 CMA patients without casein sensitization (non-Cas CMA group, age range: 0.3-6.0 years, mean 3.2 years) and 14 CMA patients with casein sensitization (Cas-CMA group, age range: 0.1-9.1 years, mean 4.5 years). The diagnosis of CMA was based on convincing clinical history, usually occurring immediately or starting several hours after the intake of moderate to large amounts of milk or its products or infant formula. Another 10 atopic controls (AC group, age range: 1.0-8.6 years, mean 4.8 years) were enrolled in the present study, among of them there were 5 children with a physician-diagnosed food allergy but no history of CMA.
Gene expression of T-related transcription factors
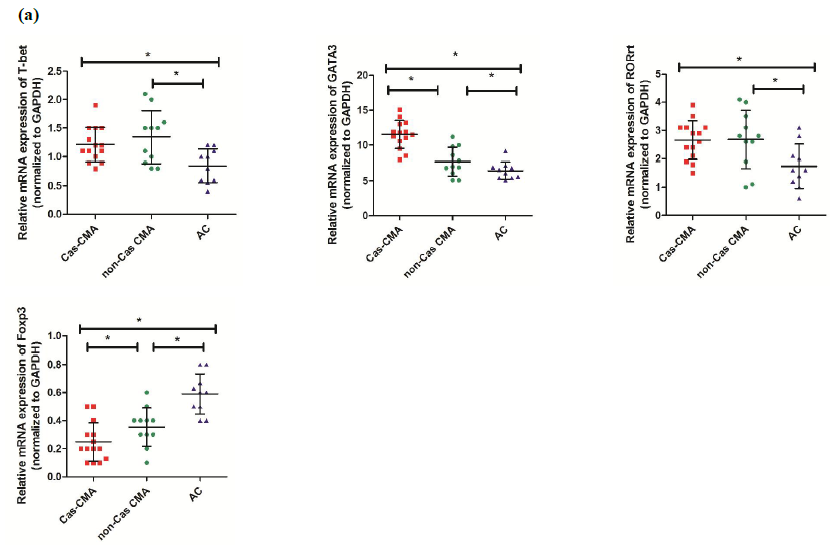
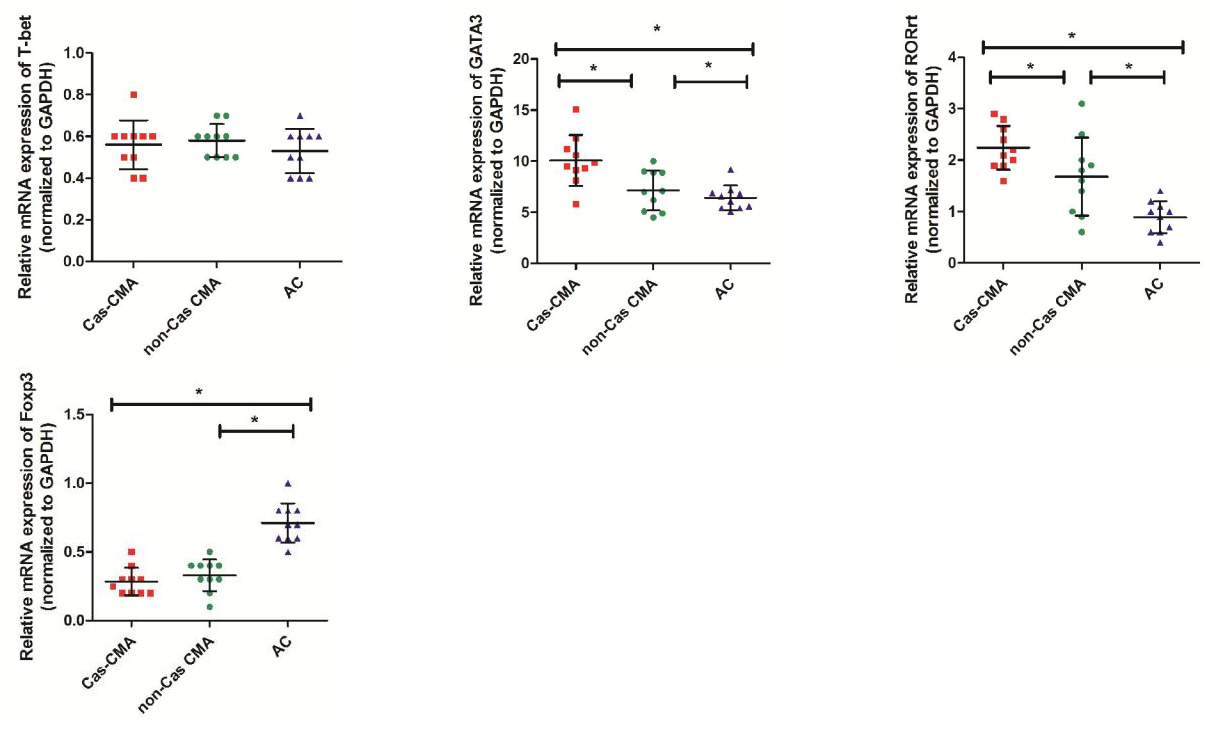
Using quantitative real-time assay, expression level of genes encoding transcription factors of Th1(T-bet), Th2(GATA3), Th17(ROR?t) and Treg(Foxp3) subsets were compared among CMA children with casein sensitization (Cas-CMA), CMA children without casein sensitization (non-Cas CMA), and atopic controls (AC). Our results showed that there was a intra-group differences existing in GATA3 or Foxp3 expression respectively, showing that a higher level of GATA3 expression (11.53±2.00 vs 7.67±2.10, P<0.05) and a decreased level of Foxp3 (0.25±0.14 vs 0.35±0.14, P<0.05) were detected in casein-sensitized CMA patients in comparison with CMA patients without casein sensitization (Figure 1a). Next, in order to describe the changing trend of T-related transcription factor among different subgroups more precisely, we isolated the total CD4+ cell population from peripheral blood and examine the expression profiles of T-related transcription factors. However, our results showed that the intra-group differences were only found in GATA3 expression (10.09±2.49 vs 7.16±1.97, P<0.05), and ROR?t (2.24±0.42 vs 1.68±0.76, P<0.05), respectively. There is a slightly decreased level of FoxP3 expression (0.29±0.10 vs 0.33±0.12, P>0.05) in CD4+T cells isolated from casein-sensitized CMA patients than those without it, this difference was not statistically significant (Figure 1b).
Th1/Th2/Th17/Treg cytokines measurement
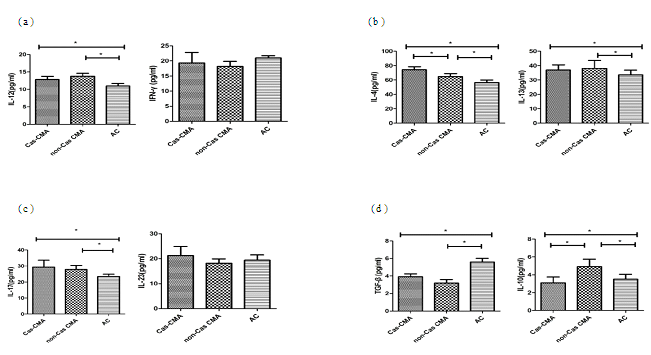
T-cell subpopulation related cytokines in patient’s serum were measured using a multiplex Luminex® detection system, enabling to detect subtle variations in a small-volume sample with a higher sensitivity. Our results showed that the average concentrations of Th1 cytokine (IL-12:13.3±3.3 pg/ml vs 11.0±2.3 pg/ml, P<0.05), Th2-related cytokines (IL-4:69.5±14.7pg/ml vs 56.5±11.2pg/ml, P<0.05; IL-13: 37.5±15.9pg/ml vs 33.6±10.2pg/ml, P<0.05), and Th17-related cytokines (IL-17: 28.6±12.3pg/ml vs 23.4±5.0pg/ml, P<0.05) were significantly increased in CMA patients than that in atopic controls. However, an obviously decreased expression of Treg-related cytokines were noted in CMA children than atopic control subjects (IL-10:4.0±2.6pg/ml vs 3.5±1.8 pg/ml, P<0.05; TGF-β: 3.6±1.3pg/ml vs 5.6±1.3pg/ml, P<0.05). Furthermore, through a intra-group comparison among CMA patients with or without casein sensitization, there showed an increase in IL-4 (74.1±16.5pg/ml vs 64.9±12.8 pg/ml, P<0.05) with a decrease in IL-10 (3.1±2.4 pg/ml vs 4.9±2.8 pg/ml, P<0.05) production in casein-sensitized CMA patient’s plasma than those patients without casein sensitization (Figure 2).
Cytokine levels of (a)Th1 (IL-12, IFN-γ), (b)Th2 (IL-4, IL-13), (c)Th17 (IL-17, IL-23) and (d)Treg (IL-10 and TGF-β) in peripheral blood from enrolled patient’s blood were detected using a custom designed MILLIPLEX MAP kit (Millipore Corporation, Bilerica, MA, USA) and the Luminex xMAP 200 System (Luminex,Austin, Texas, USA). Each symbol represents individual samples and horizontal lines show the median values. *P value <0.05 considered significant.
T cells proliferation and cytokines release after in-vitro stimulation
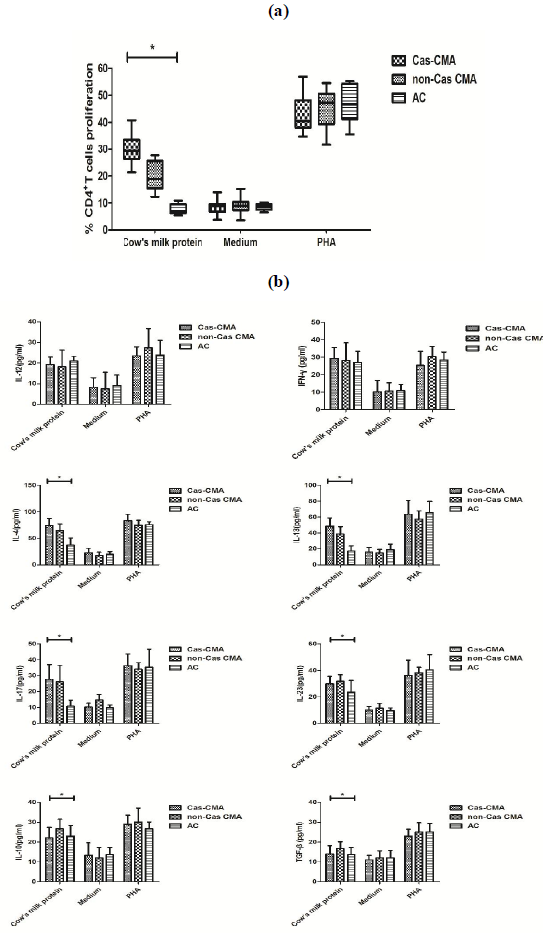
In order to explore the potential differences in cow’s milk-specific T cell reactivity, PBMCs were isolated from study subjects and stimulated with a commercial cow’s milk protein extract for 7 days in vitro. The proliferation status of cow milk-specific CD4+ T-cell frequency was assayed by CFSE dilution, showing that a significantly higher proliferation rate of cow’s milk-specific CD4+T cells in casein-sensitized CMA patients (30.0±5.8%) than that of non-Cas CMA group(20.7±5.5%) and atopic controls(7.7±1.9%) (Figure3a). In addition, upon in-vitro stimulation with cow’s milk protein, although there were significant differences in Th2 (IL-4, IL-13), Th17 (IL-17, IL-23) and Treg (IL-10 and TGF-β) among these 3 subgroups, a intra-group comparison revealed that the variations in levels of IL-4 (4.1±13.6 vs 64.7±12.3 pg/ml, P<0.05), IL-13(48.7±10.3 vs 38.7±9.4 pg/ml, P<0.05) and IL-10
(22.1±5.3 vs 26.8±4.8 pg/ml, P<0.05) between casein-sensitized CMA patients and those without casein sensitization remained statistically different (Figure 3b).
Note: PBMCs were isolated from heparinized venous blood by density-gradient separation. PBMCs (2×106 cells/mL) were cultured in vitro for 7 days at 37 °C in 5% CO2 with medium alone as negative controls, or 200 μg/mL mixture of purified alpha-, beta-, and kappa-caseins, or with 4 μg/mL phytohemagglutinin (PHA) as positive control.
- T cell proliferation rate were measured by a CFSE-based flow cytometry method.
- The release of T-related cytokines were determined by a custom designed MILLIPLEX MAP kit ( Millipore Corporation, Bilerica, MA, USA). *P value <0.05 considered significant.
Discussion
Cow’s milk is the first source of antigens encountered in infancy. The overall prognosis of cow’s milk allergy is favorable showing that the majority of infants with CMA could outgrow their allergy, however, there was still a small percentage of children retaining their persistent sensitivity to cow’s milk into the second decade [16]. Casein is the most most abundant protein components in cow’s milk, approximately accounting for 80% of total milk proteins. A cross-sectional observation study in Taiwanese population revealed that the frequency of sensitization to casein was found up to 40% of their study population [17], while were similar to those obtained form western studies [18,19]. Moreover, due to its heat stability and unique epitope structure casein protein is thought to be the most potently allergenic proteins in cow’s milk, with an evidence showing that heating milk at 120 ? for 15 min did not get rid of the antigenicity of bovine casein absolutely [20]. In real-world clinical practice, children with a persistent high casein-specific IgE are prone to exhibit a more severe phenotype of CMA and a delay of intolerance to cow’s milk [21]. Comparing with sIgE to whole cow’s milk, monitoring the level of casein-sIgE can serve as a bio-marker to predict clinical reactivity to cow’s milk with a higher sensitivity and specificity [22].
The aim of the present study was to compare the difference in T cells-mediated responses among CMA patients with or without casein sensitization. Our results showed that T cell isolated from casein-sensitized CMA patients polarized a Th2-type cytokines profiles, which featured by higher serum levels of IL-4 and IL-13 in comparison with those CMA patient without casein sensitization or atopic controls. Our results is also in line with prior studies demonstrated that casein-specific T cell clones derived from persistent CMA patients present a Th2-high signature with a higher release of IL-4 and IL-5, however, T cell responses isolated from non-allergic subjects polarized a Th1-type response [23]. Accordingly, our finding suggested that an altered T cell-mediated immunologic responses towards different protein components in cow’s milk, especially for casein allergen component, might be one of the determinant factors of the outcome for cow’s milk allergy.
Another Foxp3+regulatory T subsets (Tregs) with suppressive function were analyzed in this study. We firstly examined whether there were differences in expression level of Foxp3 among study subjects, a key transcription factor for Tregs differentiation. Our results showed that the expression levels of Foxp3 mRNA in peripheral blood mononuclear cells (PBMCs) from patients with cow’s milk allergy were significantly decreased than that in atopic controls, moreover, the variation in Foxp3 gene expression in casein-sensitized CMA patients were also found statistically different from those CMA patients without casein sensitization. Although we observed a slightly decreased level of FoxP3 expression in CD4+T cells isolated from CMA patients with casein sensitization when comparing with those without casein sensitization, the intra-group difference between these 2 groups was not statistically significant. However, the opposite results were also obtained showing an increased number of CD4+Foxp3+Tregs was found in both cow’s milk allergy group and cow’s milk tolerant group after in-vitro stimulation with casein [24]. We assumed one possible reason for this discrepancy might be explained by the different clinical features of enrolled CMA subjects. Instead of basing on OFC as enrollment criteria in other previous studies, the diagnosis of CMA in present study was mainly based on the convincing clinical history after cow’s milk consumption, the presentation of clinical symptom might include not only gastrointestinal symptoms, but also any of following urticaria, angioedema, wheezing, or a combination of above. Therefore, the differences in clinical characteristic of enrolled subjects might be one of possible reasons for the variations of Treg-mediated responses.
Next, the level of Treg-related cytokines in serum, such as IL-10 and TGF-β, were compared among different subgroups as well. Our data showed that when comparing with CMA patients without casein sensitization there was an obviously reduced serum level of IL-10 in casein-sensitized CMA patients. In addition, after in-vitro stimulation with commercial cow’s milk protein mixture, a similar result was obtained showing that the level of IL-10 was statistically lower in CMA patients with casein sensitization than those without it. IL-10 is a potent anti-inflammatory cytokine produced mainly by monocytes, lymphocytes and other cells, including macrophages, B cells, mast cells, and eosinophils [25]. IL-10 are thought to have played an important roles in maintaining intestinal homeostasis and oral tolerance induction, based on the mechanism of oral tolerance involves the deletion or anergy of T cells reactive with specific antigens associated with the expanding population of IL-10- producing regulatory T cells (Tr1) [26]. Moreover, there were much evidence reporting some single nucleotide polymorpshims (SNPs) located in IL-10 gene promoter were associated with the food allergy in Taiwanese or Japanese population, indicating that the potentially critical role of IL-10 in the pathogenesis of food allergy [27]. Moreover, Scott-Taylor et al. [28] reported that children with FA also had reduced numbers of IL-10-producing T cells in the circulation, moreover, the proportions of T cells of food sensitized patients spontaneously secreting IL-10 without antigen stimulation were lower than non-atopic controls, which was consistent with our present study.
In conclusion, the current study implicates a distinctive T cell mediated immune response among CMA patient with different sensitized component. However,inevitably, there are still some limitations in present study design. First, we should conduct standardized OFC as the enrollment criteria of CMA patients. Second, a longitudinal monitoring of casein-specific T cell response with growth status of CMA children should be further conducted, which could be helpful to provide more evidence to understand the molecular mechanism of casein-specific T cells during the CMA process.
Acknowledgement
This work was supported by a grant from Beijing Municipal Administration of Hospitals Incubating Program (No PX2018049).
Conflict of Interest Statement
We declare that we have no conflict of interest.
References
- Høst A, Halken S. A prospective study of cow milk allergy in Danish infants during the first 3 years of life. Clinical course in relation to clinical and immunological type of hypersensitivity reaction. Allergy 45 (1990): 587-596.
- Bock SA. Prospective appraisal of complaints of adverse reactions to foods in children during the first 3 years of life. Pediatrics 79 (1987) :683-688.
- Uncuoglu A, Cogurlu MT, Eser Simsek I, Ergul N, Baydemir C, Aydogan M. Predicting outgrowth of IgE-mediated cow's milk allergy: Diagnostic tests in children under two years of age. Allergol Immunopathol 2019: S0301-S0546.
- Schrander JJ, van den Bogart JP, Forget PP, Schrander-Stumpel CT, Kuijten RH, Kester AD. Cow's milk protein intolerance in infants under 1 year of age: a prospective epidemiological study. Eur J Pediatr 152 (1993): 640-644.
- Boyce JA, Assa’ad A, BurksAW, et al. Guidelines for the diagnosis and management of food allergy in the United States: summary of the NIAID-sponsored expert panel report. J Allergy Clin Immunol 126 (2010): 1105-1118.
- Santoro A, Andreozzi L, Ricci G, Mastrorilli C, Caffarelli C. Allergic reactions to cow's milk proteins in medications in childhood. Acta Biomed 90 (2019): 91-93.
- Hide DW, Guyer BM. Cows milk intolerance in Isle of Wight infants. Br J Clin Pract 37 (1983): 285-287.
- Fiocchi A, Dahdah L, Albarini M, Martelli A. Cow's milk allergy in children and adults. Chem Immunol Allergy 101 (2015): 114-123.
- Dupont C, Bradatan E, Soulaines P, Nocerino R, Berni-Canani R.Tolerance and growth in children with cow's milk allergy fed a thickened extensively hydrolyzed casein-based formula. BMC Pediatr 16 (2016): 96.
- Li J, Zhang J, Qiong C, She T, Bian Y, Lin S, Li H. Component resolved diagnostic study of cow's milk allergy in infants and young children in northern China. Int Immunopharmacol 61 (2018): 126-131.
- Cuomo B, Indirli GC, Bianchi A, Arasi S, Caimmi D, Dondi A, La Grutta S, Panetta V, Verga MC, Calvani M. Specific IgE and skin prick tests to diagnose allergy to fresh and baked cow's milk according to age: a systematic review. Ital J Pediatr 43 (2017): 93.
- Chatchatee P, Järvinen KM, Bardina L, Vila L, Beyer K, Sampson HA.Identification of IgE and IgG binding epitopes on beta- and kappa-casein in cow's milk allergic patients. Clin Exp Allergy 31 (2001): 1256-1262.
- Matsumoto N, Okochi M, Matsushima M, Kato R, Takase T, Yoshida Y, Kawase M, Isobe K, Kawabe T, Honda H. Peptide array-based analysis of the specific IgE and IgG4 in cow's milk allergens and its use in allergy evaluation. Peptides 30 (2009): 1840-1847.
- Tiemessen MM, Van Ieperen-Van Dijk AG, Bruijnzeel-Koomen CA, Garssen J, Knol EF, Van Hoffen E. Cow's milk-specific T-cell reactivity of children with and without persistent cow's milk allergy: key role for IL-10. J Allergy Clin Immunol 113 (2004): 932-939.
- Turcanu V, Maleki SJ, Lack G. Characterization of lymphocyte responses to peanuts in normal children, peanut-allergic children, and allergic children who acquired tolerance to peanuts. J Clin Invest 111 (2003): 1065-1072.
- Chen FM, Lee JH, Yang YH, Lin YT, Wang LC, Yu HH, Chiang BL. Analysis of α-lactalbumin-, β-lactoglobulin-, and casein-specific IgE among children with atopic diseases in a tertiary medical center in Northern Taiwan.J Microbiol Immunol Infect 47 (2014): 130-136.
- Restani P, Ballabio C, Di Lorenzo C, Tripodi S, Fiocchi A. Molecular aspects of milk allergens and their role in clinical events. Anal Bioanal Chem 395 (2009): 47-56.
- García-Ara MC, Boyano-Martínez MT, Díaz-Pena JM, Martín-Muñoz MF, Martín-Esteban M. Cow's milk-specific immunoglobulin E levels as predictors of clinical reactivity in the follow-up of the cow's milk allergy infants. Clin Exp Allergy 34 (2004): 866-870
- Kuitunen M, Englund H, Remes S, Movérare R, Pelkonen A, Borres MP, Mäkelä MJ. High IgE levels to α-lactalbumin, β-lactoglobulin and casein predict less successful cow's milk oral immunotherapy. Allergy 70 (2015): 955-962.
- Fiocchi A, Restani P, Riva E, Mirri GP, Santini I, Bernardo L, Galli CL. Heat treatment modifies the allergenicity of beef and bovine serum albumin.Allergy 53 (1998): 798-7802
- Babajimopoulos M, Mikolajcik EM. Influence of heat treatment on antigenicity of bovine serum albumin in milk and model systems. J Dairy Sci 61 (1978): 1380-1383.
- Caubet JC, Nowak-W?grzyn A, Moshier E, Godbold J, Wang J, Sampson HA. Utility of casein-specific IgE levels in predicting reactivity to baked milk. J Allergy Clin Immunol 131 (2013): 222-224.
- Sommanus S, Kerddonfak S, Kamchaisatian W, Vilaiyuk S, Sasisakulporn C, Teawsomboonkit W, Benjaponpitak S. Cow's milk protein allergy: immunological response in children with cow's milk protein tolerance. Asian Pac J Allergy Immunol 32 (2014): 171-177.
- Noh J, Noh G, Kim HS, Kim AR, Choi WS. Allergen-specific responses of CD19(+)CD5(+)Foxp3(+) regulatory B cells (Bregs) and CD4(+)Foxp3(+) regulatory T cell (Tregs) in immune tolerance of cow milk allergy of late eczematous reactions. Cell Immunol 274 (2012): 109-114.
- Crittenden RG, Bennett LE. Cow's milk allergy: a complex disorder. J Am Coll Nutr 24 (2005): 582S-591S.
- Yao Y, Vent-Schmidt J, McGeough MD, Wong M, Hoffman HM, Steiner TS, Levings MK. Tr1 Cells, but Not Foxp3+ Regulatory T Cells, Suppress NLRP3 Inflammasome Activation via an IL-10-Dependent Mechanism. J Immunol 195 (2015): 488-497.
- Chen TK, Lee JH, Yu HH, Yang YH, Wang LC, Lin YT, Chiang BL. Association between human IL-10 gene polymorphisms and serum IL-10 level in patients with food allergy. J Formos Med Assoc 111 (2012): 686-692.
- Scott-Taylor TH, Hourihane JB, Harper J, Strobel S. Patterns of food allergen-specific cytokine production by T lymphocytes of children with multiple allergies. Clin Exp Allergy 35 (2005): 1473-1480.