Moderate Production of Biofilm by Clinical isolates of E. coli and Pseudomonas spp in Burkina Faso
Article Information
Albert Oueremi#, 1, Abdoul Karim Ouattara#, *, 1, 3, Amana Mètuor Dabiré1, 4, Rahimatou Yasmine Wendkuni Tiemtoré1, Serge Sougué1, Jacques Simporé1, 2
1Laboratoire de Biologie Moléculaire et de Génétique (LABIOGENE), UFR-SVT, Université Joseph KI-ZERBO, 03 BP 7021 Ouagadougou 03, Burkina Faso
2Centre de Recherche Biomoléculaire Pietro Annigoni (CERBA), 01 BP 364 Ouagadougou 01, Burkina Faso
3Université Norbert Zongo, Centre Universitaire de Manga, B.P. 376 Koudougou, Burkina Faso
4Université de Dédougou, BP 176 Dédougou, Dédougou, Burkina Faso
#These authors contributed equally to this work
*Corresponding author: Abdoul Karim Ouattara, Laboratoire de Biologie Moléculaire et de Génétique (LABIOGENE), UFR-SVT, Université Joseph KI-ZERBO, 03 BP 7021 Ouagadougou 03, Burkina Faso
Received: 22 July 2023; Accepted: 01 August 2023; Published: 31 August 2023
Citation: Albert Oueremi, Abdoul Karim Ouattara, Amana Mètuor Dabiré, Rahimatou Yasmine Wendkuni Tiemtoré, Serge Sougué, Jacques Simporé. Moderate Production of Biofilm by Clinical isolates of E. coli and Pseudomonas spp in Burkina Faso. Fortune Journal of Health Sciences. 6 (2023): 300 - 305
Share at FacebookAbstract
Background: Infections caused by biofilm-producing microbes are associated with common human illnesses that are difficult to treat due to antibiotic resistance, especially when the bacteria also produce beta-lactamases. The aim of the present study was to evaluate the ability of clinical strains of Escherichia coli and Pseudomonas spp that produce beta-lactamases to produce biofilm.
Methods: The study involved two clinical strains of Escherichia coli and Pseudomonas spp that produce beta-lactamases and were isolated from pus samples at the Centre Hospitalier Universitaire Pédiatrique Charles de Gaulle of Ouagadougou, Burkina Faso. Biofilm production was assessed using the microtiter plate-based crystal violet assay, with the PA01 WT strain used as a positive control for biofilm production. Biofilm was quantified by measuring optical densities with a spectrophotometer.
Results: The E. coli strain was resistant to cefotaxime, ceftriaxone, and ceftazidime, while in addition to these antibiotics, Pseudomonas spp was resistant to imipenem. Both strains were beta-lactamase producers confirmed by the detection of blaNDM and blaIMP genes in E. coli and blaNDM, blaCTX and blaSHV in Pseudomonas spp. Optical density measurements after crystal violet staining showed that both strains were moderate biofilm producers.
Conclusions: This study highlights that clinical isolates of E. coli and Pseudomonas spp, which are responsible for human infections and produce beta-lactamases, are also moderate biofilm producers. This is a real public health concern requiring surveillance efforts and investigations to prevent and effectively combat this form of resistance.
Keywords
Biofilms, E. coli, Pseudomonas spp, β-lactamases, Burkina Faso
Biofilms articles, E. coli articles, Pseudomonas spp articles, ?-lactamases articles, Burkina Faso articles
Article Details
1. Background
Present in all ecological niches, bacteria have been the subject of extensive research worldwide since their discovery [1]. Initially studied as planktonic forms (free-swimming), the knowledge gained from these studies has greatly contributed to understanding the basic physiological processes of these microorganisms [2]. Later, work on the interactions between microorganisms and surfaces in the late 1960s and early 1970s showed that the majority of microbial biomass is attached to surfaces (biotic and abiotic) and constitutes embedded microbial populations within a matrix rich in water, sugars, and proteins, known as biofilms [2-4]. Thus, bacteria have two modes of growth: the free planktonic mode and the sessile or biofilm mode. Biofilms are one of the most widespread and successful modes of life on earth [4, 5]. Most of scientists agree that biofilms are the preferred mode of life for bacteria in the natural world, while the planktonic phase would only be a temporary stage allowing for the dissemination to new surfaces [5]. The majority of bacterial species studied in the laboratory form biofilms [6]. This mode of development confers many protections to bacteria against various environmental stressors [5] and it is often associated with public health problems, such as the formation of biofilms containing opportunistic bacteria in water networks, on medical devices (catheters, endoscopes, etc.), and on the skin and mucous membranes of the human body [7]. Once the biofilm is formed, it is very difficult to eliminate it due to its high resistance to antibacterial agents [8].
The combination of structural and physico-chemical characteristics of the biofilm gives the bacteria composing it specific properties of morphology, growth, communication between cells, and resistance to biocides [3]. Over the past few years, it has become apparent that the importance of biofilms in the medical environment is crucial, as 65% of bacterial infections are due to biofilms and over 80% of chronic bacterial infections are associated with the presence of biofilms [9]. Biofilms can form on catheters or implants (heart valves, artificial hips), and attack bodily tissues such as teeth, eyes, lungs, ears, or the urinary tract [10]. Among the pathogenic bacteria that have become real problems for human health, the gram-negative bacilli are generally found [11]. These gram-negative bacilli produce β-lactamases which are enzymes responsible for their resistance to antibiotics belonging to the β-lactam family and can form biofilms [12-14]. So, in addition to bacterial resistance through beta-lactamase production, the ability to produce biofilm confers another form of strong resistance to bacteria, leading to a complete failure of antibiotic therapy. In the present study, we report moderate biofilm production by clinical strains of Escherichia coli and Pseudomonas spp producing beta-lactamases isolated in Burkina Faso.
2. Methods
2.1 Bacterial strains
The biological material consisted of a strain of Escherichia coli and a strain of Pseudomonas spp., of human origin, isolated between 2009 and 2013 from pus samples at the Centre Hospitalier Universitaire Pédiatrique Charles De Gaulle (CHUP-CDG) of Ouagadougou, Burkina Faso [15, 16]. The isolates were identified using an API 20 E system (Bio-Mérieux, Marcy-l'Étoile, France) [17]. The strains were resistant to at least one of the cephalosporins.
2.2 Biofilm Production
The capacity for biofilm production of the isolates was assessed using the Crystal Violet staining method with some modifications. The bacteria were grown on Luria–Bertani (LB) medium and incubated at 37° C for 24 hours. The bacterial cultures were then centrifuged at 3200 rpm at 24 °C for 5 minutes and the supernatant was discarded. The bacterial pellet was washed twice (02) with 5 ml of LB and the bacteria were suspended in 1 ml of LB, and the mixture was well homogenized [18]. The bacterial suspensions were then incubated in a 96-well plate (Falcon® 96-well Polystyrene Microplates) after adjusting the DO600 nm between 0.14 et 0.16 by making dilutions. Thus, 200 μl of the bacterial preparation was distributed into the wells of the plate in triplicate and incubated without agitation at 37 °C for 24 hours. The LB medium without bacterial suspension was used as a negative control while the PA01 WT strain was used as a positive control.
After 24 hours of incubation, the culture medium and planktonic bacteria were gently removed using a micropipette. Then, the adhered bacteria were gently rinsed three times with 300 μl of water, to avoid damaging the biofilm. The bacteria were then fixed with 200 μl of methanol. After 15 minutes, the methanol was removed, and the fixed bacteria were dried in air. To reveal the biofilm, 200 μl of crystal violet was added to the wells for 15 minutes. The crystal violet was then removed, and the plates were rinsed with water to remove the excess crystal violet. The dry microplate allowed for the visual observation of the biofilm colored by the crystal violet dye and the intensity of the coloration indicated the level of biofilm formed, which could be strong, moderate, or weak. The absence of coloration indicated that no biofilm was produced. The absorbance at 590 nm was measured using the μQuant Universal Microplate Spectrophotometer (Biotek, Germany). It should be noted that 300 μl of acetic acid was used to dissolve the biofilm, and 200 μl of the mixture was then transferred to a 96-well plate for determination of the Optical Density (OD). The measured OD was then compared to the OD of the negative control (ODnc) to classify the strains as strong producers (4 × ODnc < OD), moderate producers (2 × ODnc < OD < 4 × ODnc), weak producer (ODnc < OD < 2 × ODnc) or as non-producers of biofilm (DO < DOnc) [19].
3. Results
3.1 Resistance profile
The two bacterial strains studied showed resistance to at least one beta-lactam [15-17]. The antibiogram showed that, unlike Pseudomonas spp, the E. coli strain was sensitive to imipenem (Table 1). It noteworthy that three beta-lactam resistance genes were identified in the Pseudomonas spp strain compared to two in E. coli (Table 2).
Table 1 : Antibiogram tests
|
Strains |
CTX |
CRO |
CAZ |
IMP |
|
E. coli |
R |
R |
R |
S |
|
Pseudomonas spp |
R |
R |
R |
R |
R: resistant S: sensitive, CTX: cefotaxime, CRO: ceftriaxone, CAZ: ceftazidime
Table 2 : Strain resistance genes
|
Strains |
BlaNDM |
BlaIMP |
BlaVIM |
BlaCTX |
BlaSHV |
|
Pseudomonas spp |
+ |
- |
- |
+ |
+ |
|
E. coli |
+ |
+ |
- |
- |
- |
+: presence of gene; -: absence of gene; BlaNDM: New Delhi metallo-β-lactamase; BlaIMP: Imipenemase; BlaVIM: Verona Integron-encoded Metallo-β-lactamase; BlaCTX: Cefotaximase; BlaSHV: Shylphydil Variable
3.2 Biofilm formation on Microplates
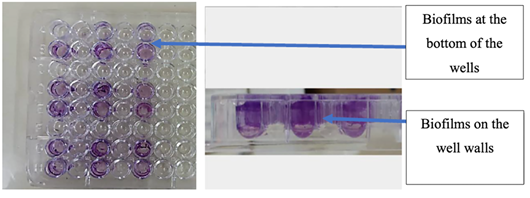
The studied strains were biofilm producers compared to the PA01 WT strain. After staining with crystal violet, the biofilms formed by the attached bacteria were visible at the bottom of the wells and on the walls of the microplate (Figure 1). The analysis of the average optical densities allowed to classify the level of biofilm formed for each strain.
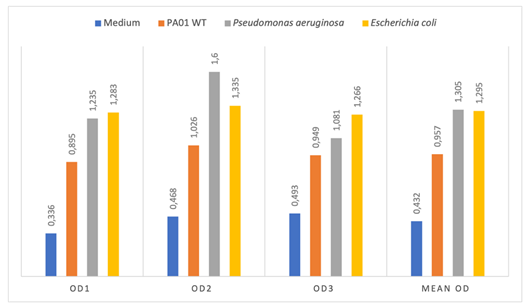
Thus, the average OD were 0.432; 0.956; 1.305; 1.295 respectively for the LB medium without inoculum (negative control); PA01 WT (positive control); Pseudomonas spp and E. coli (Figure 2). The OD of Pseudomonas spp (1.305) and E. coli (1.295) were higher than 2 x ODnc (0,864) and lower than 4 x ODnc (1,728); therefore, the studied strains were moderate biofilm producers.
4. Discussion
Gram-negative bacilli are the most involved bacteria in human infections and have developed various mechanisms of resistance [20]. In addition to known resistance mechanisms, some bacteria have the ability to produce biofilms that protect them against any attack [6]. The present study aimed to determine the ability of E. coli and Pseudomonas spp, Gram-negative bacilli, to produce biofilms in addition to their resistance through the production of beta-lactamases.
The antibiogram showed that the E. coli strain was resistant to cefotaxime (CTX), ceftriaxone (CRO), and ceftazidime (CAZ) and carried the resistance genes blaNDM and blaIMP. New Delhi metallo-β-lactamase (NDM) is a metallo-β-lactamase capable of hydrolyzing almost all beta-lactams [21]. The NDM-positive strains are generally resistant to most antimicrobial agents in addition to β-lactams due to the coexistence of other resistance mechanisms and are a global public health problem [22]. The use of carbapenem antibiotics in the treatment of severe Gram-negative bacterial infections is threatened by the emergence of the blaIMP gene among pathogenic bacteria. Previous research from our team has highlighted several bacteria strains producing extended-spectrum β-lactamases [15-17]. The Pseudomonas spp strain was resistant to the antibiotics CTX, CRO, and CAZ in addition to imipenem and carried the blaNDM, blaCTX, and blaSHV genes. These resistance genes, which are typically carried on plasmids, confer multi-resistance to the carrying bacteria, which requires new therapeutic options. Previous studies by our research team have shown that BLSE production was associated with a high co-resistance to fluoroquinolones (93% for ciprofloxacin), aminoglycosides (76.36% for gentamicin), and trimethoprim/sulfamethoxazole (95.65%) [23-25].
In this study, both the strains of E. coli and Pseudomonas spp were found to be producers of biofilm. The visualization of the plates after crystal violet staining showed that the bacteria adhered to the walls and bottoms of the wells in the microplate. The crystal violet staining assay is one of the best techniques for the detection and quantification of in vitro biofilms [26]. Our results are similar to those found by Oli et al.[27], who showed that the crystal violet staining assay is the most reliable method for detecting biofilm formation in clinical strains. The study by Oncel et al.[28] revealed that 60% of P. aeruginosa isolates from chronic rhinosinusitis produce bacterial biofilms using the crystal violet staining assay. According to a study on Gram-negative bacteria isolated from urine, the main biofilm-producing isolates were Escherichia coli strains (32.9%) followed by Pseudomonas spp strains (27.8%) (Shanmugam et al., 2017). Cepas et al.[29] found that 30.3% of E. coli isolates and 76.5% of P. aeruginosa isolates studied formed biofilms. These results confirm the ability of Pseudomonas spp and Escherichia coli isolates to form biofilms.
The biofilm quantification test showed that the clinical strains in the present study were moderate biofilm producers. A 68% (50/74) production of biofilm by clinical P. aeruginosa isolates with 4% moderate biofilm producers has been reported in the literature [30]. Similarly, Hassan et al.[31] in a study involving 110 bacterial isolates found 22.72% to be strong biofilm producers versus 40.9% to be moderate biofilm producers. Biofilm formation does not occur under the same conditions. Depending on the culture medium, temperature, and incubation time, the same bacterium can produce a weak, moderate, or strong biofilm [32]. In addition to these conditions, there may be a correlation between biofilm production and beta-lactamase production. Biofilm production could be associated with resistance enzyme production, such as beta-lactamases. The clinical isolates in the present study were both β-lactamase producers and biofilm producers. A study conducted in China showed that clinical K. pneumoniae isolates carrying blaNDM had a stronger biofilm-forming ability than K. pneumoniae carrying blaKPC [33]. However, Nirwati et al. (2019) did not find a significant association between MDR K. pneumoniae clinical strains and the ability to produce biofilm [34]. Although these two mechanisms of resistance are independent, there are enzymes present in the biofilm matrix, such as beta-lactamases, that contribute to the resistance of biofilm bacteria to antibiotics. Intensified efforts are needed to prevent the spread of multidrug-resistant bacteria in low-income countries such as Burkina Faso and the development of new endogenous therapeutic solutions to eliminate these multidrug-resistant pathogens [35-37].
5. Conclusion
The results of the present study have highlighted the moderate production of biofilm by clinical strains of Pseudomonas spp and Escherichia coli, which produce beta-lactamases. In addition to resistance through β-lactamase production, biofilm production is a preferred means of resistance for bacteria in difficult living conditions. Intensified surveillance efforts are necessary to prevent the spread of such multidrug-resistant bacteria, which pose a serious public health concern.
Declarations
Acknowledgments
The authors wish to thank all participants in this study. A deep gratitude to all the staff of Saint Camille Hospital of Ouagadougou (HOSCO) and Biomolecular Research Center Pietro Annigoni (CERBA) for technical support.
Funding
No funding was received for this study.
Authorship Contributions
Study concept and design: AKO, AMD, RYWT and JS. Study execution and acquisition of data: AO, AKO, AMD, RYWT and SS. Statistical analysis and interpretation of data: AO, AKO, AMD. Drafting of the manuscript: AKO, AKO and AMD. Critical revision of the manuscript for important intellectual content: AO, AKO, AMD, RYWT, AMD, SS and JS. Administrative, technical, and material support: AKO, AMD, RYWT and SS. Study supervision: AKO, RYWT, AMD, and JS. The Corresponding Author declare that the manuscript has been read and approved by all named authors and that the order of authors listed in the manuscript has been approved by all of us.
Competing interests
The authors declare no conflict of interest.
References
- Bahram M, Netherway T, Frioux C, Ferretti P, Coelho LP, Geisen S, et al. Metagenomic assessment of the global diversity and distribution of bacteria and fungi. Environ Microbiol (2021): 316-326.
- Yadav KK, Mandal AK and Chakraborty R. Biology of bacterial biofilms. Biology of Plants and Microbes. (2018): 61-82.
- Yin W, Wang Y, Liu L and He J. Biofilms: The Microbial "Protective Clothing" in Extreme Environments. Int J Mol Sci (2019): 1-18.
- Flemming HC and Wingender J. The biofilm matrix. Nature Reviews Microbiology (2010): 623-633.
- Rumbaugh KP and Sauer K. Biofilm dispersion. Nature Reviews Microbiology (2020): 571-586.
- Del Pozo JL. Biofilm-related disease. Expert Rev Anti Infect Ther (2018): 51-65.
- Mirzaei R, Mohammadzadeh R, Alikhani MY, Shokri Moghadam M, Karampoor S, Kazemi S, et al. The biofilm-associated bacterial infections unrelated to indwelling devices. IUBMB Life (2020): 1271-1285.
- Ciofu O, Rojo-Molinero E, Macià MD and Oliver A. Antibiotic treatment of biofilm infections. Apmis (2017): 304-319.
- Bjarnsholt T, Buhlin K, Dufrêne YF, Gomelsky M, Moroni A, Ramstedt M, et al. Biofilm formation - what we can learn from recent developments. J Intern Med (2018): 332-345.
- Rather MA, Gupta K, Bardhan P, Borah M, Sarkar A, Eldiehy KS, et al. Microbial biofilm: A matter of grave concern for human health and food industry. Journal of Basic Microbiology (2021): 380-395.
- Ruhal R and Kataria R. Biofilm patterns in gram-positive and gram-negative bacteria. Microbiological Research (2021): 126829.
- Tiemtoré RYW, Mètuor Dabiré A, Ouermi D, Sougué S, Benao S and Simporé, J. Isolation and Identification of Escherichia coli and Klebsiella pneumoniae Strains Resistant to the Oxyimino-Cephalosporins and the Monobactam by Production of GES Type Extended Spectrum Bêta-Lactamase (ESBL) at Saint Camille Hospital Center in Ouagadougou, Burkina Faso. Infect Drug Resist (2022): 3191-3204.
- Salah FD, Soubeiga ST, Ouattara AK, Sadji AY, Metuor-Dabire A, Obiri-Yeboah D, et al. Distribution of quinolone resistance gene (qnr) in ESBL-producing Escherichia coli and Klebsiella spp. in Lomé, Togo. Antimicrob Resist Infect Control (2019): 104.
- Tiemtoré RY, Mètuor-Dabiré A, Zohoncon TM, Bangré YA, Sougue S, Zongo J, et al. First Detection of PE-Type Extended-Spectrum β-lactamases at Saint Camille Hospital Center of Ouagadougou, Burkina Faso. Int. J. Biochem. Biophys. Mol Biol (2019): 7-12.
- Metuor Dabire A, Zongo KJ, Zeba B, Moussawi J, Baucher M and El Jaziri M. Resistances to the Oxyimino-Cephalosporins by Ctx-M-15 Producing Klebsiella Isolated from the Urines Samples of Patients in the University Hospital Complex Paediatric Charles De Gaulle (CHUP-CDG) Of Ouagadougou in Burkina Faso Journal of Asian Scientific Research (2013): 882-890.
- Metuor Dabiré A, Tiemtoré RYW-K, Bangré YA, Zohoncon T, Mahoukèdè Sougué S, Zongo KJ, et al. Detection of multidrug-resistant enterobacteria simultaneously producing extended-spectrum -lactamases of the PER and GES types isolated at Saint Camille Hospital Center, Ouagadougou, Burkina Faso. African Journal of Microbiology Research (2019): 414-420.
- Metuor Dabire A, Zongo KJ, Zeba B, Traoré/Ouedraogo R, Moussawi J, Baucher M, et al. First detection of SHV-type extended spectrum β-lactamases in the University Hospital Complex Paediatric Charles De Gaulle (CHUP-CDG) of Ouagadougou in Burkina Faso. Journal of Asian Scientific Research (2014): 214-221.
- Ramos-Vivas J, Chapartegui-González I, Fernández-Martínez M, González-Rico C, Fortún J, Escudero, R, et al. Biofilm formation by multidrug resistant Enterobacteriaceae strains isolated from solid organ transplant recipients. Scientific Reports (2019): 8928.
- Hassan A, Usman J, Kaleem F, Omair M, Khalid A, and Iqbal M. Evaluation of different detection methods of biofilm formation in the clinical isolates. Brazilian Journal of Infectious Diseases (2011): 305-311.
- Breijyeh Z, Jubeh B and Karaman, R. Resistance of Gram-Negative Bacteria to Current Antibacterial Agents and Approaches to Resolve It. Molecules (2020).
- Wu W, Feng Y, Tang G, Qiao F, McNally A, and Zong Z. NDM Metallo-β-Lactamases and Their Bacterial Producers in Health Care Settings. Clin Microbiol Rev (2019): 1-45.
- Dortet, L, Poirel, L, and Nordmann, P. Worldwide dissemination of the NDM-type carbapenemases in Gram-negative bacteria. Biomed Res Int (2014): 249856.
- Moumouni A, Diagbouga S, Nadembèga C, Metuor Dabire A, Ouattara K, and Zohoncon T. Quinolone Resistance (qnr) genes in fecal carriage of extended Spectrum beta-lactamases producing Enterobacteria isolated from children in Niger. Curr Res Microbiol Biotechnol (2017): 953-7.
- Diagbouga S, Salah F, Sadji A, Dabire A, Nadembega C, Kere A, et al. Detection of High Prevalence of TEM/SHV CTX-M Genes in ESBL Producing and Multidrug Resistant Klebsiella pneumoniae and Klebsiella oxytoca. J Clin Diagn Res (2016): 000129.
- Salah F, Diagbouga S, Dabire AM, Sadji A, Nadembega C, and Moumouni A. First detection of Resistance genes encoding extended Spectrum β-lactamase producing Escherichia coli at Lomé, Togo. Archives of Clinical Microbiology (2016): 1-7.
- Bellifa, S, Hassaine, H, Damien, B, Nicolas, C, Imane, M, Ibtissem, K, et al. Evaluation of biofilm formation of Klebsiella pneumoniae isolated from medical devices at the University Hospital of Tlemcen, Algeria. African Journal of Microbiology Research (2013): 5558-5564.
- Oli AK, Raju S, Nagaveni S, and Chandrakanth RK. Biofilm formation by multidrug resistant Enterococcus faecalis (MDEF) originated from clinical samples. Journal of Microbiology and Biotechnology Research (2012): 284-288.
- Oncel S, Pinar E, Sener G, Calli C and Karagoz U. Evaluation of bacterial biofilms in chronic rhinosinusitis. J Otolaryngol Head Neck Surg (2010): 52-5.
- Cepas V, López Y, Muñoz E, Rolo D, Ardanuy C, Martí S, et al. Relationship Between Biofilm Formation and Antimicrobial Resistance in Gram-Negative Bacteria. Microbial Drug Resistance (2019): 72-79.
- Lima JLdC, Alves LR, Paz dJNP, Rabelo MA, Maciel MAV and Morais dMMC. Analysis of biofilm production by clinical isolates of <i>Pseudomonas aeruginosa</i> from patients with ventilator-associated pneumonia. Revista Brasileira de Terapia Intensiva (2017): 310-316.
- Hassan H and Hanna S. Extended-spectrum beta-lactamases (ESBLs) detection in some uropathogenic bacteria and their correlation with biofilm formation. Zanco Journal of Medical Sciences (2019): 375-382.
- Tolker-Nielsen, T. Biofilm Development. Microbiol Spectr (2015): Mb-0001-2014.
- Duan Q, Wang Q, Sun S, Cui Q, Ding Q, Wang R, et al. ST11 Carbapenem-Resistant Klebsiella pneumoniae Clone Harboring bla(NDM) Replaced a bla(KPC) Clone in a Tertiary Hospital in China. Antibiotics (Basel) (2022): 1373.
- Nirwati H, Sinanjung K, Fahrunissa F, Wijaya F, Napitupulu S, Hati VP et al. Biofilm formation and antibiotic resistance of Klebsiella pneumoniae isolated from clinical samples in a tertiary care hospital, Klaten, Indonesia. BMC Proceedings (2019): 1-8.
- Rouamba A, Ouédraogo V, Compaoré E, Compaoré M and Kiendrebeogo M. Free Radical Scavenging Capacity and Anti-Biofilm Potentiality of Six Wild Edible Fruits from Burkina Faso. International Journal of Current Microbiology and Applied Sciences (2023): 2085-2093.
- Ouedraogo V, Rouamba A, Compaoré E, Compaoré M, and Kiendrebeogo M. Antioxidant, antiquorum-sensing and antibiofilm activities of Balanites aegyptiaca (L.) Del.(Balanitaceae) and Terminalia macroptera Guill. and Perr. (Combretaceae). Adv. Biochem (2018): 26-31.
- Issa K, Vincent O, Ablassé R, Moussa C and Martin K. Anti-quorum sensing and anti-biofilm activities of Securidaca longepedunculata Fresen, an endangered species from Burkina Faso. African Journal of Microbiology Research (2021): 146-151.