Microbiota Metabolites Affecting Host Diseases and Welfare
Article Information
Annie Fang1, Igor F Tsigelny1,2,3, Valentina L Kouznetsova2*
1REHS program, San Diego Supercomputer Center, UC San Diego, La Jolla, CA, 92093, USA
2San Diego Supercomputer Center, UC San Diego, La Jolla, CA, 92093, USA
3Department of Neurosciences, UC San Diego, La Jolla, CA, 92093, USA
*Corresponding Author: Valentina L Kouznetsova, San Diego Supercomputer Center, UC San Diego, La Jolla, CA, 92093, USA
Received: 28 December 2020; Accepted: 07 January 2021; Published: 11 January 2021
Citation: Annie Fang, Igor F Tsigelny, Valentina L Kouznetsova. Microbiota Metabolites Affecting Host Diseases and Welfare. Fortune Journal of Health Sciences 4 (2021): 191-206.
Share at FacebookAbstract
The microbiome is a community of over 100 trillion micro-organisms living together in a particular habitat. Recently, because many studies have shown a significant correlation between the microbiome and the host’s health and pathogenesis of diseases like Type 2 Diabetes (T2D), research on the microbiome has surged in popularity. Special attention has been paid to the microbiome present in the gastrointestinal tract because it is the largest, most diverse, and controlling of digestion. In this review, we analyze the benefits of a healthy microbiome and the causes and effects of microbiome dysbiosis. Furthermore, we elucidate how lifestyle factors like diet and exercise affect the microbiome’s microbial composition, its production of metabolites, and their relationship with disease. In this review, we will summarize the current knowledge in this field.
Keywords
Microbiota; Microbiome; Microbial composition; Metabolites; Type 2 Diabetes; T2D
Microbiota articles; Microbiome articles; Microbial composition articles; Metabolites articles; Type 2 Diabetes articles; T2D articles, Microbiota articles Microbiota Research articles Microbiota review articles Microbiota PubMed articles Microbiota PubMed Central articles Microbiota 2023 articles Microbiota 2024 articles Microbiota Scopus articles Microbiota impact factor journals Microbiota Scopus journals Microbiota PubMed journals Microbiota medical journals Microbiota free journals Microbiota best journals Microbiota top journals Microbiota free medical journals Microbiota famous journals Microbiota Google Scholar indexed journals Microbiome articles Microbiome Research articles Microbiome review articles Microbiome PubMed articles Microbiome PubMed Central articles Microbiome 2023 articles Microbiome 2024 articles Microbiome Scopus articles Microbiome impact factor journals Microbiome Scopus journals Microbiome PubMed journals Microbiome medical journals Microbiome free journals Microbiome best journals Microbiome top journals Microbiome free medical journals Microbiome famous journals Microbiome Google Scholar indexed journals Microbial composition articles Microbial composition Research articles Microbial composition review articles Microbial composition PubMed articles Microbial composition PubMed Central articles Microbial composition 2023 articles Microbial composition 2024 articles Microbial composition Scopus articles Microbial composition impact factor journals Microbial composition Scopus journals Microbial composition PubMed journals Microbial composition medical journals Microbial composition free journals Microbial composition best journals Microbial composition top journals Microbial composition free medical journals Microbial composition famous journals Microbial composition Google Scholar indexed journals Metabolites articles Metabolites Research articles Metabolites review articles Metabolites PubMed articles Metabolites PubMed Central articles Metabolites 2023 articles Metabolites 2024 articles Metabolites Scopus articles Metabolites impact factor journals Metabolites Scopus journals Metabolites PubMed journals Metabolites medical journals Metabolites free journals Metabolites best journals Metabolites top journals Metabolites free medical journals Metabolites famous journals Metabolites Google Scholar indexed journals Type 2 Diabetes articles Type 2 Diabetes Research articles Type 2 Diabetes review articles Type 2 Diabetes PubMed articles Type 2 Diabetes PubMed Central articles Type 2 Diabetes 2023 articles Type 2 Diabetes 2024 articles Type 2 Diabetes Scopus articles Type 2 Diabetes impact factor journals Type 2 Diabetes Scopus journals Type 2 Diabetes PubMed journals Type 2 Diabetes medical journals Type 2 Diabetes free journals Type 2 Diabetes best journals Type 2 Diabetes top journals Type 2 Diabetes free medical journals Type 2 Diabetes famous journals Type 2 Diabetes Google Scholar indexed journals T2D articles T2D Research articles T2D review articles T2D PubMed articles T2D PubMed Central articles T2D 2023 articles T2D 2024 articles T2D Scopus articles T2D impact factor journals T2D Scopus journals T2D PubMed journals T2D medical journals T2D free journals T2D best journals T2D top journals T2D free medical journals T2D famous journals T2D Google Scholar indexed journals deoxyribonucleic acid articles deoxyribonucleic acid Research articles deoxyribonucleic acid review articles deoxyribonucleic acid PubMed articles deoxyribonucleic acid PubMed Central articles deoxyribonucleic acid 2023 articles deoxyribonucleic acid 2024 articles deoxyribonucleic acid Scopus articles deoxyribonucleic acid impact factor journals deoxyribonucleic acid Scopus journals deoxyribonucleic acid PubMed journals deoxyribonucleic acid medical journals deoxyribonucleic acid free journals deoxyribonucleic acid best journals deoxyribonucleic acid top journals deoxyribonucleic acid free medical journals deoxyribonucleic acid famous journals deoxyribonucleic acid Google Scholar indexed journals toll-like receptor articles toll-like receptor Research articles toll-like receptor review articles toll-like receptor PubMed articles toll-like receptor PubMed Central articles toll-like receptor 2023 articles toll-like receptor 2024 articles toll-like receptor Scopus articles toll-like receptor impact factor journals toll-like receptor Scopus journals toll-like receptor PubMed journals toll-like receptor medical journals toll-like receptor free journals toll-like receptor best journals toll-like receptor top journals toll-like receptor free medical journals toll-like receptor famous journals toll-like receptor Google Scholar indexed journals Ruminococcus torques articles Ruminococcus torques Research articles Ruminococcus torques review articles Ruminococcus torques PubMed articles Ruminococcus torques PubMed Central articles Ruminococcus torques 2023 articles Ruminococcus torques 2024 articles Ruminococcus torques Scopus articles Ruminococcus torques impact factor journals Ruminococcus torques Scopus journals Ruminococcus torques PubMed journals Ruminococcus torques medical journals Ruminococcus torques free journals Ruminococcus torques best journals Ruminococcus torques top journals Ruminococcus torques free medical journals Ruminococcus torques famous journals Ruminococcus torques Google Scholar indexed journals
Article Details
1. Introduction
The microbiome is a community of over 100 trillion micro-organisms including bacteria, fungi, protozoa, and viruses that live together in a particular habitat. This habitat can be located at many different sites of the human body such as the skin, mouth, lungs, and gastrointestinal tract, and their composition can vary greatly. “Microbiome” and “microbiota” are words that are commonly used interchangeably, but their meanings differ completely. "Microbiota" is a community of microorganisms in and on the host body. “Microbiome”refers to the genetic material of all the microbes, such as deoxyribonucleic acid (DNA) and ribonucleic acid (RNA). On the other hand, “microbiota” refers to only the microbes in a particular habitat. Both are hypothesized to influence characteristics of their residing habitats and as a result, the host’s health.
2. Methods
We analyzed scholarly articles from PubMed and Google Scholar with key search terms of “Microbiome causes Type 2 Diabetes”, “Microbiome and Disease”, “Microbiome”, and “Microbiota Metabolites”. It was our goal to understand and summarize what is currently known about the microbiota and its relationship with the host’s wellbeing. However, as the microbiota is so diverse with over 100 trillion microorganisms, it is poorly understood and there are many studies with conflicting results.
3. Results
3.1 Mutual Symbiosis of the Healthy Microbiota and Host
It is widely hypothesized that a healthy microbiota and the host can have a mutually symbiotic relationship. A healthy microbiota could play a role in strengthening the gut’s integrity or shaping the intestinal epithelium (the lining of the small and large intestines of the gastrointestinal tract), as well as in harvesting energy by producing vitamins and short-chain fatty acids (SCFAs), anti-proliferative lipids that represent an essential energy source for the body [1]. It can also protect against pathogens which is largely believed to be possible because of interactions between microbial organisms and pathogens, such as competition for the same nutrients. This would in turn be regulating host immunity. Additionally, the microbiota modulates digestion of dietary fiber as well as the host’s energy balance [2].
3.2 Microbiota Dysbiosis
But while a healthy microbiota may have many benefits, dysbiosis of the microbiota, which is mainly defined by an imbalance of microbes, might be very harmful. Many studies, such as one done by Thursby and Juge, seem to show that one effect of dysbiosis is an increased risk in pathogenesis of diseases like cancer, inflammatory bowel disease (IBD), and metabolic disease [1]. Obesity and malnutrition are common effects of microbial dysbiosis, and this might be because the imbalance of microbes could lead to increased or decreased efficiency at yielding energy from the diet.
Some authors suggest that dysbiosis can cause the “leaky gut” syndrome [3]. Essentially, scientists hypothesize that the human tissue always contains low amounts of bacteria. Then, as an effect of inflammation, disruptions to physical and molecular barriers, and immunosuppression, which can all be caused by dysbiosis, the barriers of the human tissue could be weakened, allowing bacteria to pass into otherwise sterile areas. For instance, Frasinariu and colleagues found that the gut-derived bacterial products—lipopolysaccharides (LPS) and unmethylated CpG DNA—were able to permeate the lining of the gut and activate toll-like receptor (TLR) signaling pathway in patients with non-alcoholic fatty liver disease (weakened barriers) [4]. As TLR affects activation of NFkB, which is related to inflammation, the “leakiness” of the gut may be a cause of microbiome dysbiosis [3].
3.3 Microbial Composition
Special attention has been paid to the microbial composition of the microbiome because it may affect gut phenotypes and thus have a significant impact on one’s health. There is a lot of research that has been conducted to elucidate the relationship between the two main phyla: Bacteroidetes and Firmicutes. Though there has been some conflicting results, the most well accepted conclusion is that increased levels of Firmicutes and decreased levels of Bacteroidetes play a role in microbial dysbiosis and often lead to obesity. For example, according to Koliada with co-authors, an increase in Firmicutes as well as a decrease in Bacteroidetes correlates to obesity [5]. Additionally, in a study conducted by Ley and colleagues, it was found that genetically obese mice had 50% less Bacteroidetes and 50% more Firmicutes in comparison to lean mice despite similar diets and activity levels [6].
Eckburg and a team also saw an increased ratio of Firmicutes to Bacteroidetes in IBD, but furthermore saw increased Ruminococcus torques and E. coli in Crohn’s disease, and lower levels of Bifidobacteria and small-intestinal bacterial overgrowth in chronic fatigue syndrome [8].
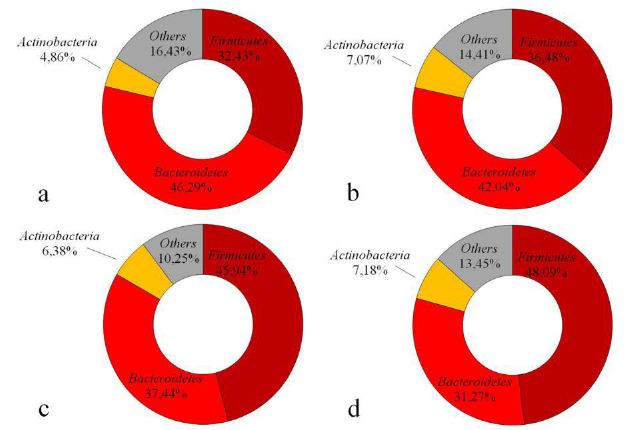
Figure 1: The microbial compositions for hosts of different BMI categories (a BMI < 18.5, b BMI 18.5–24.9, c BMI 25–29.9 and d BMI ≥ 30). As the BMI increases, the levels of Firmicutes increases and those of Bacteroidetes decreases. Reproduced from the open access article [5].
3.4 Influence of Diet on Microbial Composition
It is believed that one factor that shapes the microbiota’s composition is the host’s diet. This is likely because the diet defines the host’s total energy intake and macronutrient composition; in essence it is the sustenance for the growth of gut bacteria. One influence of the diet is, as mentioned above, its impact on microbial diversity. But more specifically, Mosca and co-authors suggested that diet can influence the level of bacterial predators inside the intestinal microbiota [9]. The “western” diet, which is characterized by foods high in fat and sugar such as fast food, may be associated with a loss of bacterial predators such as protists, bacteriophages, and predatory bacteria [9]. This is harmful because stable ecosystems require competition and cooperation, something that can only be achieved through a balance of prey and predators [10]. The “western” diet has also been associated with decreased levels of Bifidobacteria, Bacteroidetes, and Prevotella phyla, but also an increase in the Firmicutes phylum [11,12].
Furthermore, the intake of specific nutrients may alter microbial compositions. It was shown that increasing protein intake led to increased propionate and Bacteroidetes in the microbiome [13-15]. But on the other hand, studies conducted by Duncan and colleagues as well as Walker and co-authers that diets that were low in fiber and carbohydrates led to reduced butyrate and Firmicutes, as Firmicutes produce butyrate [16,17]. High-fiber diets have also been shown to enhance the gut microbiota diversity and improve glucose control [18].
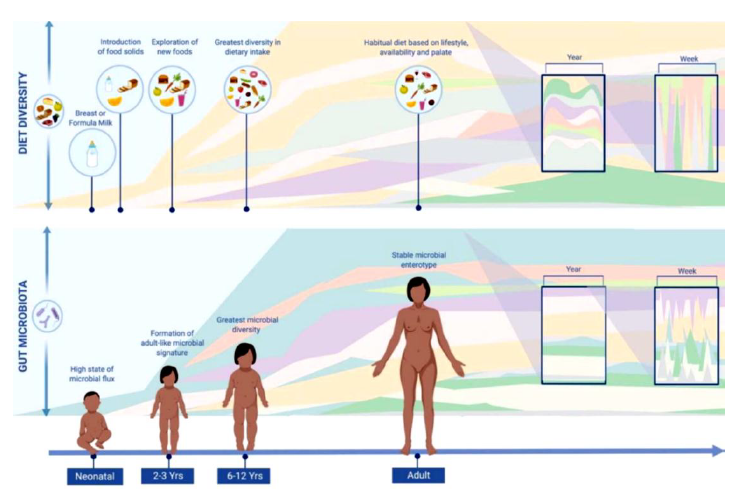
Figure 2: Diversity in dietary intake is directly correlated to microbial diversity. Reproduced from the open access article [11].
The abovementioned findings show that diet plays a role in microbial composition, but some researchers, such as David and co-authors, suggested that diet not only causes significant alteration of microbes, but does it rapidly [19]. In their study, David and colleagues, provided humans with two diets: a “plant-based diet” full of grains, legumes, fruits, and vegetables; and an “animal-based diet” full of meats, eggs, and cheeses [19]. It was found that the microbial composition of both groups was altered significantly in only one day after consuming the given diet and when they returned to their original diets, their microbial compositions reverted to their original structures in only two days. Authors hypothesized that the microbiome’s rapid adaptation correlates to human evolution in which our ancestors had to be flexible to both herbivorous and carnivorous diets because the availability of food was dependent on the season or their hunting success [19].
However, the relationship between diet and microbial composition is not perfectly understood. Zinöcker and Lindseth reminded that despite many studies showing links between the “western” diet and disease, the biological mechanisms that explain how this works is unknown [20]. They suggested that the “western” diet indirectly affects microbial composition by promoting inflammation, which would then in turn affect the microbiome [20]. Also a study by Chassaing and co-authors argued that it is not necessarily the fat in a high-fat “western” diet that leads to inflammation in the gut but rather the lack of plant-derived nutrients such as fiber and phytochemicals [21].
3.5 Influence of Exercise on Microbial Composition
Another factor that potentially shapes microbial composition is exercise. According to a study done by Schranner and colleagues, it was shown that levels of lactate, pyruvate, TCA cycle intermediates, fatty acids, acylcarnitines, and ketone bodies all usually increased after exercise while levels of bile acids decreased [22]. However, it must be noted that this study does not state if they are measuring metabolites indigenous to the host or produced by the microbiome. Furthermore, Mitchell and colleagues reported that exercise is associated with increased population of butyrate producing bacteria and thus also fecal butyrate [23]. But these researchers admitted that there was an unclear risk of bias in this study and thus there must be more research on the relationship between exercise and microbial composition.
3.6 Microbiota Metabolites
The microbiota, like the human body, can also produce, modulate, and degrade many metabolites (Table 1). According to Fischbach and Sonnenburg and to Nicholson with co-authors, there are instances when some metabolic processes can only be completed by the microbiota, such as the degradation of certain complex proteins and carbohydrates [24,25]. It is important to remember that there is a difference in metabolites associated with bacterial metabolism and metabolites produced as a product of microbial–host cometabolism. Bacterial or microbial metabolism is defined as the process in which microbes obtain the energy and nutrients necessary for their survival. Meanwhile, microbial–host cometabolism is the process in which the degradation of a compound requires the presence of other primary substrates. Metabolites produced from this process are also commonly referred to as bacterial transformed compounds.
3.7 Bacterial Metabolism
Table 1: Summary of metabolites produced in bacterial metabolism
|
Metabolites |
Bacteria |
References |
|
Short-chain fatty acids (SCFAs): acetic acid, butyric acid, propionic acid |
Bacteroides, Firmicutes |
[26,27] |
|
Organic acids: benzoate, hippurate, phenylacetate, phenylpropionate, cresol, hydroxybenzoate, hydroxyphenylacetate, hydroxyphenyl propionate, 3,4-dihydroxyphenyl propionate, tricarballylate, D-lactate, D-arabinitol |
Faecalibacterium prausnitzii, Bifidobacterium |
[28,29] |
|
Vitamins: thiamine, folate, biotin, riboflavin, pantothenic acid, vitamin K, vitamin B12 |
Bacteroides, Bifidobacterium, Enterococcus |
[30,31] |
Some of the most well-known metabolites that result from bacterial metabolism are SCFAs, but their effects on the host are debated. The most abundant SCFAs: acetate, propionate, and butyrate exist in the microbial community at an average ratio of 3:1:1 respectively, although it can differ substantially based on diet and microbial composition [32].
There has been much discussion about the role of SCFAs for the host. Scheppach and co-authors suggest that acetate can raise the mucosal blood flow which helps protect against harmful agents [33]. They add that propionate can inhibit cholesterol synthesis and thus decrease the risk of high cholesterol and the cardiovascular problems that follow. Furthermore, butyrate has been shown to have many beneficial effects such as improving insulin sensitivity, nourishing the colonic mucosa, and being a substantial energy source for colonocytes, or epithelial cells of the colon [34]. Samuel and colleagues also argue that SCFAs can activate G-protein-coupled receptors like GPR41 and GPR43, which are related to inflammation and enteroendocrine regulation [35].
But there is also research that supports the contrary, or that SCFAs and vitamin production may be related to obesity and metabolic syndromes. For instance, Vernocchi and co-authors say that an abundance of butyrate can lead to promotion of certain cancers [36]. The contradicting opinions on the role of SCFAs proves that this topic still needs more research. The abundance of SCFAs is suggested to be used as a biomarker for one’s health [37]. This conclusion is supported by Turnbaugh with colleagues, Meyer and Hostetter, and Schwiertz with a team, who found that total fecal SCFA concentration increased with obese patients and decreased after anti-obesity treatment [7,38,39]. As Turnbaugh and co-authors compared which plasma solutes were less prevalent in the germ-free and conventional rats used in their study with high-resolution mass spectrometry, we can agree that the identified metabolites were of microbial origin with relative confidence [7].
3.8 Bacterial Transformed Compounds
Microbiota transforms a set of host compounds in microbial–host co-metabolism (Table 2).
Table 2: Summary of the bacterial transformed compounds produced by microbes in microbial–host co-metabolism
|
Metabolites |
Bacteria |
References |
|
Bile salts: chocolate, chenodeoxycholate, deoxycholate, lithocholate |
Lactobacillus, Enterococcus, Bifidobacterium, Clostridium, Bacteroides |
[40-43] |
|
Polyphenols: flavonoids, hydroxycinnamic acids |
Bacteroides, Lactobacillus, Enterococcus, Bifidocaterium, Ruminococcus, Escherichia coli |
[45,46] |
|
Lipids: glycerol |
Clostridium, Lactobacillus |
[47,48] |
|
Amino acids |
Clostridium botulinum, Clostridium sporogenes |
[49,50] |
It has been established that the microbiome will transform bile acids [44], lipids, polyphenols, and amino acids through microbial–host cometabolism [36]. Oliphant and Allen-Vercoe suggest that it is harder to incorporate the 20 amino acid building blocks of amino acids than to incorporate monosaccharide units into biochemical pathways [51]. Thus, microbes will typically not ferment amino acids and instead transform them to other compounds because monosaccharides like carbohydrates are more efficient to use as energy. We will summarize the significant bacterial transformed compounds elucidated from Oliphant's and Allen-Vercoe’s studies [51]. Bifidobacterium (Actinobacteria) is transformed to acetate, ethanol, formate, and lactate. Bacteroides (Bacteroidetes) is transformed to 1,2-propanediol, carbon dioxide and hydrogen, ethanol, formate, propionate, and succinate. Clostridium (Firmicutes) is transformed to 1,2-propanediol, acetate, carbon dioxide and hydrogen, ethanol, formate, lactate, propionate, butyrate, and valerate. Escherichia (Proteobacteria) is transformed to 1,2-propanediol, 2,3-butanediol, acetate, carbon dioxide and hydrogen, ethanol, formate, lactate, and succinate [51]. In addition, Wu with co-authors suggest that microbes can transform tryptophan to indoxyl sulfate and indolepropionic acid, and carnitine and choline to trimethylamine, which can then by oxidized into trimethylamine N oxide (TMAO) [52].
3.9 Microbiota Metabolites and Disease
As more and more research links microbial dysbiosis to disease, there has been increasingly more attention paid to the metabolites that are produced either by bacterial metabolism or microbial–host cometabolism. In one study by Amedei and co-authors, it is suggested that metabolites derived from the gut microbiota can impact the circulatory system in two ways [53]. Microbes can first stimulate the enteric nervous system, which would in turn affect the brain centers that modulate the cardiovascular system. They could affect tissues and organs that control homeostasis of the circulatory system such as the heart and blood cells [53]. Thus, an imbalance in microbial metabolites caused by dysbiosis could potentially lead to pathogenesis of disease.
Some metabolites that have been linked to disease are: TMAO, uremic toxins, and peptides. Several studies have found that TMAO activates mitogen activated protein kinase (MAPK) and NF-kB signaling, which are related to pathogenesis of diseases [54]. It has been reported that high levels of TMAO can also serve as a prediction of serious cardiovascular problems such as myocardial infarction, stroke, or even death [55].
Uremic toxins are produced by the metabolization of amino acids such as tyrosine, phenylalanine, and tryptophan in the microbiome. Some toxins include indoxyl sulfate, indoxyl glucuronide, indoleacetic acid, p-cresyl sulfate, p-cresyl glucuronide, phenyl sulfate, phenyl glucuronide, phenylacetic acid, and hippuric acid [54]. Some of these toxins have been identified as predictors of disease but the exact way this is done is not well understood. For instance, Hsu with a team demonstrate that indoxyl sulfate is associated with coronary atherosclerosis and Itoh with colleagues also found increased uremix toxins in hemodialysis patients [56,57]. Additionally, the release of peptides could worsen intestinal inflammation and disrupt the gut–blood barrier, which could allow bacteria to pass into the circulatory system [53].
According to Chang and co-authors, butyrate, one of the most prevalent SCFAs in the microbiome, activates the IL6 gene [58]. As IL6 has been shown to be involved in the development of inflammation, insulin resistance, and β-cell dysfunction, activation of IL6 would be increasing the risk of these symptoms [59]. Chang's and co-authors' study [58] can be used reliably because the microbes observed in the study were grown directly in a laboratory setting rather than taken from the body so there is no confusion between microbial metabolites and metabolites indigenous to the host. Furthermore, butyrate as well as propionate have been shown to induce activity of NF-kB [60]. Because excessive activation of NF-kB may lead to insulin resistance, metabolites that activate NF-kB would be increasing the risk of disease [61].
3.10 Microbiota and T2D
Dysbiosis of the microbiota has been associated with the pathogenesis of many diseases, and its relationship with T2D is one of the well-studied. First, there is evidence that the microbial composition of the microbiome plays an important role in increased T2D risk. Many studies show relationships between certain bacteria phyla and T2D. For example, though the genera of Ruminococcus, Fusobacterium, and Blautiah have inconsistent reports of their species levels, studies have consistently shown their positive association with T2D [62,63]. On the other hand, the genera of Bifidobacterium, Bacteroides, Faecalibacterium, Akkermansia, and Roseburia were negatively associated with T2D [19]. Bifidobacterium is most commonly reported for its negative association with T2D. Almost all animal studies that were given a Bifidobacterium strain showed improved glucose tolerance, showing that Bifidobacterium could play a protective role in T2D [64]. Similar studies were conducted on mice to test the association of Bacteroides and T2D, and when Bacteroides acidifaciens and Bacteroides uniformis were administered, the mice showed improved glucose tolerance and insulin resistance [65,66]. Additionally, concentrations of Bacteroides intestinalis, Bacteroides 20-3, and Bacteroides vulgatus decreased in T2D patients [67]. These observations support that the Bacteroides genus is negatively associated with T2D. Finally, the Roseburia, Faecalibacterium, and Akkermansia genera have all been reported to have decreased concentrations in T2D patients [68,69]. Five studies have shown lower frequencies of Roseburia, or more specifically Roseburia invulinivornas and Roseburia_272, in patients with T2D in comparison to the healthy control group [62,70]. Additionally, four out of five studies have shown lower levels of Faecalibacterium in T2D patients [69,71]. The last phylum of Akkermansia is only recently discovered but scientists have seen its role in host glucose metabolism and its negative association with T2D [72]. As these studies support that different bacteria have stronger and weaker associations with T2D, we can be confident that the microbial composition of the microbiome impacts T2D risk.
Another area of importance when understanding the causes of T2D is the fiber concentrations produced by microbial organisms. Karlsson et al. argue that T2D is associated with reduced concentrations of fiber-degrading bacteria [69]. This is likely because regular consumption of dietary fiber can help prevent erosion of the intestinal mucus barrier, which in turn helps protect against invasive pathogens [73]. Dietary fiber is a major fuel source for the microbiota and it impacts the bacteria, composition, and functional activity of the microbiome. In a study done on mice, it was shown that a lack of fiber led to decreased microbiota composition and thus inflammation and insulin resistance [74]. Additionally, a study by Cani et al. demonstrated that the fiber inulin leads to higher concentrations of A. muciniphila, which is a microbe that shows reduced levels in patients with T2D [75]. While fiber seems beneficial in decreasing the risk of T2D, Slujis with colleagues demonstrates that diets with high concentrations of glucose, such as the “western” diet, increase the risk of T2D by reducing microbial diversity [76]. Scientists hypothesize that this is because they are high in digestible starches but lack sufficient quantities of fiber.
3.11 Relationship between T2D and Lifestyle Habits
In examining numbers of T2D patients based on world geography, it is further emphasized that lifestyle habits are impactful in pathogenesis of T2D. In the world, the Middle East has one of the highest percentages of T2D in adults. More specifically, the World Health Organization labels the Gulf countries: Kuwait, Bahrain, Saudi Arabia, and the United Arab Emirates, as some of the countries with the highest rates of obesity [77].
Table 3: Percentages of T2D in the countries with the highest percentages of T2D in adults (WHO estimates 2010)
|
Country |
Male |
Female |
|
Kuwait |
36% |
48% |
|
Saudi Arabia |
28% |
44% |
|
UAE |
25% |
42% |
|
Bahrain |
21% |
38% |
|
Qatar |
19% |
32% |
|
Lebanon |
15% |
27% |
|
Oman |
8% |
17% |
After the study of the lifestyle habits of these countries, it is clear that urbanization, improvements in living conditions, technological advances, and cultural restrictions are important factors to blame. People living in the cities have much higher rates of T2D than those in rural areas of the Middle East, and this is attributed to more sedentary city life and an abundance of unhealthy food options like fast-food chains. Meanwhile, rural life is far more active as people typically depend more on physically laboring work such as fishing and agriculture [78]. Additionally, because the Gulf countries have become massively wealthy as crucial oil-exporting countries, people have experienced higher incomes and are able to purchase more expensive products such as those with meat, egg, and milk [79]. It is also important that typical options at fast-food restaurants contain meat, egg, and milk. Because of their high accessibility, these products make up much of the Middle Eastern diet yet they contain little to no fiber. This contributes to weak mucosal protection from pathogens, reduced microbial diversity, an imbalance in the ratio of Firmicutes to Bacteroidetes, and decreased concentrations of beneficial SCFAs. All these factors can cause significant microbiota dysbiosis and as a result worsen the risk of T2D.
Aside from unhealthy diets, the sedentary life in cities also plays a role in the Middle East’s high numbers of T2D patients. Because of an influx of personal wealth as well as more manufacture of automobiles, the availability of automobiles like cars and buses has increased vastly. As driving or being driven requires much less physical effort than simpler forms of transport such as walking or cycling, this change significantly contributes to the people’s lack of exercise. It is also interesting that women had higher rates of obesity than men in every country (Table 3). This is likely because in addition to the unhealthy diets and dependency on automobiles for transport, there are also many cultural restrictions of women, such as limits to sports and exercise. The lack of exercise that is associated with Middle Eastern cities could lead to decreased butyrate production and worsened insulin resistance. This would in turn increase the risk of T2D.
4. Discussion
Our research concludes that microbial composition and the metabolites produced by either bacterial metabolism or microbial–host cometabolism can be both beneficial and harmful to the host. A healthy microbiome could have a mutually symbiotic relationship with the host by producing metabolites such as SCFAs, and dietary fibers that have been shown to not only provide energy but also protect against disease. On the other hand, an imbalance in the microbiome’s composition often leads to microbial dysbiosis which weakens the host’s immune system and thus worsens the risk of T2D pathogenesis. Often an imbalance in composition is an alteration in the ratio of the Bacteroidetes and Firmicutes phyla. An increase in Firmicutes and a decrease in Bacteroidetes has been associated with microbial dysbiosis [5].
It is clear that metabolite production and microbial composition is likely synonymous with the host’s health, so the factors that influence production of metabolites, such as diet and exercise, play an ever-important role in maintaining the host’s health. Results from many studies conclude that diets high in fiber may improve gut diversity and glucose control while “western diets”, or diets high in fat and sugar, may lead to a loss of microbial diversity. Exercise is also influential in metabolite production as increased levels of exercise have been associated with increased levels of butyrate and SCFAs that can help control inflammation and prevent the “leakiness” of the gut. Daily lifestyle habits have significant effects on the microbiome and T2D. But as studies show the much higher prevalence of T2D in Middle Eastern cities in comparison to rural towns, it is possible that larger developments such as urbanization, improving living conditions, technological advances, and cultural restrictions can also affect the host’s microbiome.
These factors will affect both the metabolites directly produced from bacterial metabolism, such as SCFAs, organic acids, and vitamins, along with metabolites that are bacterially transformed such as bile salts, polyphenols, lipids, and amino acids. Production of some metabolites like TMAO, uremic toxins, and peptides, have also been linked to microbial dysbiosis and often pathogenesis of diseases like T2D. The relationship between the T2D and the microbiome is represented by the positive and negative associations that certain bacteria phyla have with pathogenesis of T2D. For instance, the phylum of Fusobacterium shows positive associations with T2D while the phyla of Bifidobacterium and Bacteroidetes phylum show negative associations with T2D.
Overall, despite the connection between the microbiota and pathogenesis of diseases such as T2D still being poorly understood, results from recent research have increasingly shown the importance of the microbiome’s composition in preventing microbial dysbiosis and improving the host’s health. With further research, the possibility of scientists being able to better understand and manipulate the microbiota for our own benefit becomes ever larger.
Conflict of Interest:
Authors declare no conflict of interest.
Funding:
No funding was received.
References
- Thursby E, Juge N. Introduction to the human gut microbiota. Biochemical Journal 474 (2017): 1823-1836.
- Rios-Covian D, Salazar N, Gueimonde M, et al. Shaping the metabolism of intestinal Bacteroides population through diet to improve human health. Frontiers in Microbiology 8 (2017): 376.
- Kawai T, Akira S. Signaling to NF-κB by Toll-like receptors. Trends in Molecular Medicine 13 (2007): 460-469.
- Frasinariu OE, Ceccarelli S, Alisi A, et al. Gut-liver axis and fibrosis in nonalcoholic fatty liver disease: an input for novel therapies. Digestive and Liver Disease 45 (2013): 543-551.
- Koliada A, Syzenko G, Moseiko V, et al. Association between body mass index and Firmicutes/Bacteroidetes ratio in an adult Ukrainian population. BMC Microbiology 17 (2017): 120.
- Ley RE, Bäckhed F, Turnbaugh P, et al. Obesity alters gut microbial ecology. Proceedings of the National Academy of Sciences 102 (2005): 11070-11075.
- Turnbaugh PJ, Ley RE, Mahowald MA, et al. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 444 (2006): 1027.
- Eckburg PB, Bik EM, Bernstein CN, et al. Diversity of the human intestinal microbial flora. Science 308 (2005): 1635-1638.
- Mosca A, Leclerc M, Hugot JP. Gut microbiota diversity and human diseases: should we reintroduce key predators in our ecosystem?. Frontiers in Microbiology 7 (2016): 455.
- Elmqvist T, Folke C, Nyström M, et al. Response diversity, ecosystem change, and resilience. Frontiers in Ecology and the Environment 1 (2003): 488-494.
- Leeming ER, Johnson AJ, Spector TD, et al. Effect of diet on the gut microbiota: Rethinking intervention duration. Nutrients 11 (2019): 2862.
- Clarke SF, Murphy EF, Nilaweera K, et al. The gut microbiota and its relationship to diet and obesity: new insights. Gut Microbes 3 (2012): 186-202.
- Furet JP, Kong LC, Tap J, et al. Differential adaptation of human gut microbiota to bariatric surgery–induced weight loss: links with metabolic and low-grade inflammation markers. Diabetes 59 (2010): 3049-3057.
- Jumpertz R, Le DS, Turnbaugh PJ, et al. Energy-balance studies reveal associations between gut microbes, caloric load, and nutrient absorption in humans. The American Journal of Clinical Nutrition 94 (2011): 58-65.
- Liou AP, Paziuk M, Luevano JM, et al. Conserved shifts in the gut microbiota due to gastric bypass reduce host weight and adiposity. Science Translational Medicine 5 (2013): 178ra41.
- Duncan SH, Belenguer A, Holtrop G, et al. Reduced dietary intake of carbohydrates by obese subjects results in decreased concentrations of butyrate and butyrate-producing bacteria in feces. Applied and Environmental Microbiology 73 (2007): 1073-1078.
- Walker AW, Ince J, Duncan SH, et al. Dominant and diet-responsive groups of bacteria within the human colonic microbiota. The ISME Journal 5 (2011): 220-230.
- Makki K, Deehan EC, Walter J, et al. The impact of dietary fiber on gut microbiota in host health and disease. Cell Host & Microbe 23 (2018): 705-715.
- David LA, Maurice CF, Carmody RN, et al. Diet rapidly and reproducibly alters the human gut microbiome. Nature 505 (2014): 559-563.
- Zinöcker MK, Lindseth IA. The Western diet–microbiome-host interaction and its role in metabolic disease. Nutrients 10 (2018): 365.
- Chassaing B, Miles-Brown J, Pellizzon M, et al. Lack of soluble fiber drives diet-induced adiposity in mice. American Journal of Physiology-Gastrointestinal and Liver Physiology 309 (2015): G528-G541.
- Schranner D, Kastenmüller G, Schönfelder M, et al. Metabolite Concentration Changes in Humans After a Bout of Exercise: a Systematic Review of Exercise Metabolomics Studies. Sports Medicine-Open 6 (2020): 11.
- Mitchell CM, Davy BM, Hulver MW, et al. Does exercise alter gut microbial composition? A systematic review. Medicine & Science in Sports & Exercise 51 (2019): 160-167.
- Fischbach MA, Sonnenburg JL. Eating for two: how metabolism establishes interspecies interactions in the gut. Cell Host & Microbe 10 (2011): 336-347.
- Nicholson JK, Holmes E, Kinross J, et al. Host-gut microbiota metabolic interactions. Science 336 (2012): 1262-1267.
- Macfarlane S, Macfarlane GT. Regulation of short-chain fatty acid production. Proceedings of the Nutrition Society 62 (2003): 67-72.
- Li Z, Quan G, Jiang X, et al. Effects of metabolites derived from gut microbiota and hosts on pathogens. Frontiers in Cellular and Infection Microbiology 8 (2018): 314.
- Lord RS, Bralley JA. Clinical applications of urinary organic acids. Part 2. Dysbiosis markers. Altern Med Rev 13 (2008): 292-306.
- Li M, Wang B, Zhang M, et al. Symbiotic gut microbes modulate human metabolic phenotypes. Proceedings of the National Academy of Sciences 105 (2008): 2117-2122.
- Martens JH, Barg H, Warren MA, et al. Microbial production of vitamin B 12. Applied Microbiology and Biotechnology 58 (2002): 275-285.
- Hill MJ. Intestinal flora and endogenous vitamin synthesis. European Journal of Cancer Prevention: the official journal of the European Cancer Prevention Organisation (ECP) 6 (1997): S43-S45.
- Martin AM, Sun EW, Rogers GB, et al. The influence of the gut microbiome on host metabolism through the regulation of gut hormone release. Frontiers in Physiology 10 (2019): 428.
- Scheppach W, Pomare EW, Elia M, et al. The contribution of the large intestine to blood acetate in man. Clinical Science 80 (1991): 177-182.
- Scheppach W. Effects of short chain fatty acids on gut morphology and function. Gut 35 (1994): S35-S38.
- Samuel BS, Shaito A, Motoike T, et al. Effects of the gut microbiota on host adiposity are modulated by the short-chain fatty-acid binding G protein-coupled receptor, Gpr41. Proceedings of the National Academy of Sciences 105 (2008): 16767-16772.
- Vernocchi P, Del Chierico F, Putignani L. Gut microbiota profiling: metabolomics based approach to unravel compounds affecting human health. Frontiers in Microbiology 7 (2016): 1144.
- Ríos-Covián D, Ruas-Madiedo P, Margolles A, et al. Intestinal short chain fatty acids and their link with diet and human health. Frontiers in Microbiology 7 (2016): 185.
- Meyer TW, Hostetter TH. Uremic solutes from colon microbes. Kidney International 81 (2012): 949-954.
- Schwiertz A, Taras D, Schäfer K, et al. Microbiota and SCFA in lean and overweight healthy subjects. Obesity 18 (2010): 190-195.
- Elkins CA, Moser SA, Savage DC. Genes encoding bile salt hydrolases and conjugated bile salt transporters in Lactobacillus johnsonii 100-100 and other Lactobacillus species. Microbiology 147 (2001): 3403-3412.
- Franz CM, Specht I, Haberer P, et al. Bile salt hydrolase activity of enterococci isolated from food: screening and quantitative determination. Journal of Food Protection 64 (2001): 725-729.
- Grill J, Schneider F, Crociani J, et al. Purification and characterization of conjugated bile salt hydrolase from Bifidobacterium longum BB536. Applied and Environmental Microbiology 61 (1995): 2577-2582.
- Rossocha M, Schultz-Heienbrok R, von Moeller H, et al. Conjugated bile acid hydrolase is a tetrameric N-terminal thiol hydrolase with specific recognition of its cholyl but not of its tauryl product. Biochemistry 44 (2005): 5739-5748.
- Urdaneta V, Casadesús J. Interactions between bacteria and bile salts in the gastrointestinal and hepatobiliary tracts. Frontiers in Medicine 4 (2017): 163.
- Duda-Chodak A, Tarko T, Satora P, et al. Interaction of dietary compounds, especially polyphenols, with the intestinal microbiota: a review. European Journal of Nutrition 54 (2015): 325-341.
- Selma MV, Espin JC, Tomas-Barberan FA. Interaction between phenolics and gut microbiota: role in human health. Journal of Agricultural and Food Chemistry 57 (2009): 6485-6501.
- Parsons JB, Rock CO. Bacterial lipids: metabolism and membrane homeostasis. Progress in Lipid Research 52 (2013): 249-276.
- Elsden SR, Hilton MG, Waller JM. The end products of the metabolism of aromatic amino acids by Clostridia. Archives of Microbiology 107 (1976): 283-288.
- Fonknechten N, Chaussonnerie S, Tricot S, et al. Clostridium sticklandii, a specialist in amino acid degradation: revisiting its metabolism through its genome sequence. BMC Genomics 11 (2010): 1-2.
- Dodd D, Spitzer MH, Van Treuren W, et al. A gut bacterial pathway metabolizes aromatic amino acids into nine circulating metabolites. Nature 551 (2017): 648-652.
- Oliphant K, Allen-Vercoe E. Macronutrient metabolism by the human gut microbiome: major fermentation by-products and their impact on host health. Microbiome 7 (2019): 91.
- Wu WK, Hsu CC, Sheen LY, et al. Measurement of gut microbial metabolites in cardiometabolic health and translational research. Rapid Communications in Mass Spectrometry 34 (2020): e8537.
- Amedei A, Morbidelli L. Circulating metabolites originating from gut microbiota control endothelial cell function. Molecules 24 (2019): 3992.
- Seldin MM, Meng Y, Qi H, et al. Trimethylamine N-oxide promotes vascular inflammation through signaling of mitogen-activated protein kinase and nuclear factor-κ Journal of the American Heart Association 5 (2016): e002767.
- Wang Z, Klipfell E, Bennett BJ, et al. Gut flora metabolism of phosphatidylcholine promotes cardiovascular disease. Nature 472 (2011): 57-63.
- Hsu CC, Lu YC, Chiu CA, et al. Levels of indoxyl sulfate are associated with severity of coronary atherosclerosis. Clinical and Investigative Medicine (2013): E42-E49.
- Itoh Y, Ezawa A, Kikuchi K, et al. Correlation between serum levels of protein-bound uremic toxins in hemodialysis patients measured by LC/MS/MS. Mass Spectrometry 2 (2013): S0017.
- Chang PV, Hao L, Offermanns S, et al. The microbial metabolite butyrate regulates intestinal macrophage function via histone deacetylase inhibition. Proceedings of the National Academy of Sciences 111 (2014): 2247-2252.
- Akbari M, Hassan-Zadeh V. IL-6 signalling pathways and the development of type 2 diabetes. Inflammopharmacology 26 (2018): 685-698.
- Lakhdari O, Tap J, Béguet-Crespel F, Le Roux K, de Wouters T, Cultrone A, Nepelska M, Lefevre F, Doré J, Blottiere HM. Identification of NF-κB modulation capabilities within human intestinal commensal bacteria. BioMed Research International 2011 (2011): 282356
- Andreasen AS, Kelly M, Berg RM, et al. Type 2 diabetes is associated with altered NF-κB DNA binding activity, JNK phosphorylation, and AMPK phosphorylation in skeletal muscle after LPS. PloS One 6 (2011): e23999.
- Candela M, Biagi E, Soverini M, et al. Modulation of gut microbiota dysbioses in type 2 diabetic patients by macrobiotic Ma-Pi 2 diet. British Journal of Nutrition 116 (2016): 80-93.
- He Y, Wu W, Zheng HM, et al. Regional variation limits applications of healthy gut microbiome reference ranges and disease models. Nature Medicine 24 (2018): 1532-1535.
- Moya-Pérez A, Neef A, Sanz Y. Bifidobacterium pseudocatenulatum CECT 7765 reduces obesity-associated inflammation by restoring the lymphocyte-macrophage balance and gut microbiota structure in high-fat diet-fed mice. PloS One 10 (2015): e0126976.
- Yang JY, Lee YS, Kim Y, et al. Gut commensal Bacteroides acidifaciens prevents obesity and improves insulin sensitivity in mice. Mucosal Immunology 10 (2017): 104-116.
- Cano PG, Santacruz A, Moya Á, et al. Bacteroides uniformis CECT 7771 ameliorates metabolic and immunological dysfunction in mice with high-fat-diet induced obesity. PloS One 7 (2012): e41079.
- Wu X, Ma C, Han L, et al. Molecular characterisation of the faecal microbiota in patients with type II diabetes. Current Microbiology 61 (2010): 69-78.
- Zhang X, Shen D, Fang Z, et al. Human gut microbiota changes reveal the progression of glucose intolerance. PloS One 8 (2013): e71108.
- Karlsson FH, Tremaroli V, Nookaew I, et al. Gut metagenome in European women with normal, impaired and diabetic glucose control. Nature 498 (2013): 99-103.
- Murphy R, Tsai P, Jüllig M, et al. Differential changes in gut microbiota after gastric bypass and sleeve gastrectomy bariatric surgery vary according to diabetes remission. Obesity Surgery 27 (2017): 917-925.
- Gao R, Zhu C, Li H, et al. Dysbiosis signatures of gut microbiota along the sequence from healthy, young patients to those with overweight and obesity. Obesity 26 (2018): 351-361.
- Greer RL, Dong X, Moraes AC, et al. Akkermansia muciniphila mediates negative effects of IFN [gamma] on glucose metabolism. Nature Communications 7 (2016): 13329.
- Desai MS, Seekatz AM, Koropatkin NM, et al. A dietary fiber-deprived gut microbiota degrades the colonic mucus barrier and enhances pathogen susceptibility. Cell 167 (2016): 1339-1353.
- Zou J, Chassaing B, Singh V, et al. Fiber-mediated nourishment of gut microbiota protects against diet-induced obesity by restoring IL-22-mediated colonic health. Cell Host & Microbe 23 (2018): 41-53.
- Cani PD. Gut microbiota and obesity: lessons from the microbiome. Briefings in Functional Genomics 12 (2013): 381-387.
- Sluijs I, Forouhi NG, Beulens JW, et al. The amount and type of dairy product intake and incident type 2 diabetes: results from the EPIC-InterAct Study. The American Journal of Clinical Nutrition 96 (2012): 382-390.
- ALNohair S. Obesity in gulf countries. Interna-tional Journal of Health Sciences (Qassim) 8 (2014): 79-83.
- Mohsen MA, Rahman IA, Al Khadra AH, et al. Prevalence of physical inactivity in Saudi Arabia: a brief review. East Mediterr Health J 10 (2004): 663-670.
- Al Othaimeen AI, Al Nozha M, Osman AK. Obesity: an emerging problem in Saudi Arabia. Analysis of data from the National Nutrition Survey. EMHJ-Eastern Mediterranean Health Journal 13 (2007): 441-448.
