Development and Qualification of a Novel Method for Quantitative Analysis of Lipid Aldehydes in Tissue
Article Information
William C. Putnam1,2,3*, Sarah M. Swenson1,3, Dinesh Rakheja4
1Department of Pharmaceutical Sciences, Jerry H. Hodge School of Pharmacy, Dallas, TX, 75235, USA
2Department of Pharmacy Practice, Jerry H. Hodge School of Pharmacy, Dallas, TX, 75235, USA
3Clinical Pharmacology and Experimental Therapeutics Center, Jerry H. Hodge School of Pharmacy, Dallas, TX, 75235, USA
4Departments of Pathology and Pediatrics, University of Texas Southwestern Medical Center, Dallas, TX, 75390, USA
*Corresponding Author: William C. Putnam, Department of Pharmaceutical Sciences, Pharmacy Practice, and Clinical Pharmacology and Experimental Therapeutics Center, Jerry H. Hodge School of Pharmacy, Dallas, TX, 75235, USA
Received: 03 November 2020; Accepted: 12 November 2020; Published: 28 November 2020
Citation:
William C. Putnam, Sarah M. Swenson, Dinesh Rakheja. Development and Qualification of a Novel Method for Quantitative Analysis of Lipid Aldehydes in Tissue. Journal of Analytical Techniques and Research 2 (2020): 123-136.
Share at FacebookAbstract
Abstract
There exists much interest in identifying and quantitating oxidized lipids in biofluids and tissues because certain oxidized lipids may potentially serve as early-stage diagnostic biomarkers of disease states such as preeclampsia. To that end, a set of seven biologically relevant oxidized lipids (lipid aldehydes) have been studied in placental tissue. A novel method for analyzing these seven target biologically relevant oxidized lipids was developed and is presented. Placental samples processed and analyzed by LC-MS/MS for the content of the seven specified lipid aldehydes. Additionally, the placental samples were profiled for additional (untargeted) lipid aldehydes. The LC-MS/MS method was qualified and found to have acceptable linearity, accuracy and precision for its intended use. It was determined that placental tissue from mothers with preeclampsia had relatively lower amounts of oxidized lipids when compared to control placental tissue.
Keywords
Preeclampsia; Biomarker; Oxidized lipid; Lipid aldehyde; LC-MS/MS
Preeclampsia articls; Biomarker articls; Oxidized lipid articls; Lipid aldehyde articls; LC-MS/MS articls
Preeclampsia articles Preeclampsia Research articles Preeclampsia review articles Preeclampsia PubMed articles Preeclampsia PubMed Central articles Preeclampsia 2023 articles Preeclampsia 2024 articles Preeclampsia Scopus articles Preeclampsia impact factor journals Preeclampsia Scopus journals Preeclampsia PubMed journals Preeclampsia medical journals Preeclampsia free journals Preeclampsia best journals Preeclampsia top journals Preeclampsia free medical journals Preeclampsia famous journals Preeclampsia Google Scholar indexed journals Biomarker articles Biomarker Research articles Biomarker review articles Biomarker PubMed articles Biomarker PubMed Central articles Biomarker 2023 articles Biomarker 2024 articles Biomarker Scopus articles Biomarker impact factor journals Biomarker Scopus journals Biomarker PubMed journals Biomarker medical journals Biomarker free journals Biomarker best journals Biomarker top journals Biomarker free medical journals Biomarker famous journals Biomarker Google Scholar indexed journals Oxidized lipid articles Oxidized lipid Research articles Oxidized lipid review articles Oxidized lipid PubMed articles Oxidized lipid PubMed Central articles Oxidized lipid 2023 articles Oxidized lipid 2024 articles Oxidized lipid Scopus articles Oxidized lipid impact factor journals Oxidized lipid Scopus journals Oxidized lipid PubMed journals Oxidized lipid medical journals Oxidized lipid free journals Oxidized lipid best journals Oxidized lipid top journals Oxidized lipid free medical journals Oxidized lipid famous journals Oxidized lipid Google Scholar indexed journals Lipid aldehyde articles Lipid aldehyde Research articles Lipid aldehyde review articles Lipid aldehyde PubMed articles Lipid aldehyde PubMed Central articles Lipid aldehyde 2023 articles Lipid aldehyde 2024 articles Lipid aldehyde Scopus articles Lipid aldehyde impact factor journals Lipid aldehyde Scopus journals Lipid aldehyde PubMed journals Lipid aldehyde medical journals Lipid aldehyde free journals Lipid aldehyde best journals Lipid aldehyde top journals Lipid aldehyde free medical journals Lipid aldehyde famous journals Lipid aldehyde Google Scholar indexed journals LC-MS/MS articles LC-MS/MS Research articles LC-MS/MS review articles LC-MS/MS PubMed articles LC-MS/MS PubMed Central articles LC-MS/MS 2023 articles LC-MS/MS 2024 articles LC-MS/MS Scopus articles LC-MS/MS impact factor journals LC-MS/MS Scopus journals LC-MS/MS PubMed journals LC-MS/MS medical journals LC-MS/MS free journals LC-MS/MS best journals LC-MS/MS top journals LC-MS/MS free medical journals LC-MS/MS famous journals LC-MS/MS Google Scholar indexed journals malondialdehyde articles malondialdehyde Research articles malondialdehyde review articles malondialdehyde PubMed articles malondialdehyde PubMed Central articles malondialdehyde 2023 articles malondialdehyde 2024 articles malondialdehyde Scopus articles malondialdehyde impact factor journals malondialdehyde Scopus journals malondialdehyde PubMed journals malondialdehyde medical journals malondialdehyde free journals malondialdehyde best journals malondialdehyde top journals malondialdehyde free medical journals malondialdehyde famous journals malondialdehyde Google Scholar indexed journals colorimetric assay articles colorimetric assay Research articles colorimetric assay review articles colorimetric assay PubMed articles colorimetric assay PubMed Central articles colorimetric assay 2023 articles colorimetric assay 2024 articles colorimetric assay Scopus articles colorimetric assay impact factor journals colorimetric assay Scopus journals colorimetric assay PubMed journals colorimetric assay medical journals colorimetric assay free journals colorimetric assay best journals colorimetric assay top journals colorimetric assay free medical journals colorimetric assay famous journals colorimetric assay Google Scholar indexed journals biological matrices articles biological matrices Research articles biological matrices review articles biological matrices PubMed articles biological matrices PubMed Central articles biological matrices 2023 articles biological matrices 2024 articles biological matrices Scopus articles biological matrices impact factor journals biological matrices Scopus journals biological matrices PubMed journals biological matrices medical journals biological matrices free journals biological matrices best journals biological matrices top journals biological matrices free medical journals biological matrices famous journals biological matrices Google Scholar indexed journals analytical strategies articles analytical strategies Research articles analytical strategies review articles analytical strategies PubMed articles analytical strategies PubMed Central articles analytical strategies 2023 articles analytical strategies 2024 articles analytical strategies Scopus articles analytical strategies impact factor journals analytical strategies Scopus journals analytical strategies PubMed journals analytical strategies medical journals analytical strategies free journals analytical strategies best journals analytical strategies top journals analytical strategies free medical journals analytical strategies famous journals analytical strategies Google Scholar indexed journals quantitate MDA articles quantitate MDA Research articles quantitate MDA review articles quantitate MDA PubMed articles quantitate MDA PubMed Central articles quantitate MDA 2023 articles quantitate MDA 2024 articles quantitate MDA Scopus articles quantitate MDA impact factor journals quantitate MDA Scopus journals quantitate MDA PubMed journals quantitate MDA medical journals quantitate MDA free journals quantitate MDA best journals quantitate MDA top journals quantitate MDA free medical journals quantitate MDA famous journals quantitate MDA Google Scholar indexed journals
Article Details
1. Introduction
Early diagnosis of preeclampsia provides the opportunity for clinicians to optimize medical management and enhance therapeutic outcomes for infants and mothers. Identification of predictive and prognostic biomarkers offers the promise of earlier management and is an area of active investigation [1-5]. Yet, no reliable predictive or prognostic biomarkers are currently used in clinical practice. The goal here is to identify predictive and prognostic biomarkers of preeclampsia.
Pregnancy related diseases can lead to medical complications that extend beyond the pregnancy for both mother and child. Some of these diseases such as preeclampsia, HELLP (hemolysis, elevated liver enzymes, low platelet count), and AFLP (acute fatty liver pregnancy) are clinical syndromes of multi-factorial origin. The final common pathway for the development of these multi-system disorders may be maternal endothelial dysfunction due to oxidative stress [6]. The placenta may play a crucial role in the production of oxidants that lead to this damage. Highlighting this theory, during preeclampsia it has been demonstrated that lipid oxidation is increased and the levels of glutathione and vitamin C are also increased. Furthermore, the activities of antioxidant enzymes superoxide dismutase, glucose 6-phosphate dehydrogenase, glutathione peroxidase, and glutathione S-transferase are decreased in preeclamptic placentas [7,8]. Reduced levels of total thiol content and superoxide dismutase, observed in the plasma of preeclamptic women, increased significantly after delivery [8]. The products of lipid oxidation may serve as potential early-stage biomarkers; therefore, investigation into the presence, quantity, and predictive nature (biomarkers) of fatty-acid oxidation products such as lipid aldehydes in placenta and other biological fluids is prudent. Furthermore, the uncovering of biomarkers in placenta may also provide a diagnostic measure for the extent of oxidative stress experienced during pregnancy. This measure of oxidative stress may then be useful in correlation with, and eventually prediction of outcomes.
Traditionally, malondialdehyde (MDA) has been the primary marker for oxidative stress, [9] and typically, the 2-thiobarbituric colorimetric assay has been extensively used for quantitation of MDA in various biological matrices [10,11]. This assay has come under scrutiny in the past few years due to its lack of specificity [10,11]. Consequently, numerous other analytical strategies have been employed to quantitate MDA, as well as other biomarkers. For example, direct infusion mass spectrometry has recently been used to quantitate lipid-based biomarkers in serum from preeclamptic women [12]. This study utilized an untargeted lipid profiling approach and demonstrated that there are global changes in the lipid profiles. Additional methods include GC/MS, HPLC with UV detection, and more recently LC-MS and LC-MS/MS [13-16].
The investigation of lipid aldehydes by LC-MS is complicated by the fact that underivatized lipid aldehydes do not produce a strong mass spectrometric response under atmospheric pressure ionization conditions. To enhance the mass spectral response, derivatization can be employed. This is highlighted by recent elegant work of Andreoli and co-workers who employed 2,4-dinitrophenyhydrazone (2,4-DNP) derivatives and LC-MS/MS to study lipid aldehydes [12]. In this paper they developed and validated a method to quantitate lipid aldehydes in breath condensate following derivatization with 2,4-DNP.
Seven key lipid aldehydes that have been implicated to be produced under oxidative stress conditions were initially selected for study here [9,12]. These include MDA, acrolein, hexanal, heptanal, nonanal, 4-hydroxyhexenal, 4-hydroxynonenal. By investigating this small targeted set of lipid-analytes there remained the possibility that a number of other potential biomarkers derived from lipid oxidation may be missed; therefore, there was also interest in developing methodology to ‘profile’ biological samples for other lipid aldehydes produced under oxidative stress conditions.
In reviewing different approaches, a second class of aldehyde derivatization reagents was found that possess properties that may render them superior to 2,4-DNP. This other class of reagents, termed Girard reagents, also employs hydrazone formation for covalent attachment (Figure 1) [17,18].
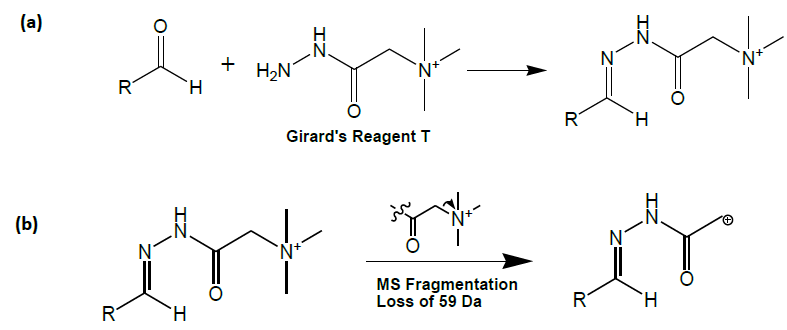
Figure 1: (a) Reaction of Girard reagent T with an aldehyde; formation of the hydrazone bond and (b) Collision induced dissociation fragmentation of a Girard reagent T conjugate of an aldehyde.
Moreover, Girard reagents provide a conjugate that is already positively charged, thus facilitating the formation of gas-phase ions. Derivatization with Girard reagents is not new to the field of biological sample analysis and mass spectrometry. David Johnson of the Women’s and Children’s Hospital in Australia has employed Girard reagents in combination with electrospray ionization tandem mass spectrometry to profile ketosteroids in the diagnosis of congenital adrenal hyperplasia [19]. Griffiths and co-workers used both Girard reagents T and P to produce conjugates of and subsequently analyze neutral oxosteroids [20,21]. Girard reagents have been used with matrix assisted laser desorption ionization mass spectrometry to analyze small oligosaccharides [22]. Lai and co-workers employed Girard reagent P in the µLC-MS/MS analysis of 17-hydroxyprogesterone from dried blood spots [23]. Girard reagents were used with secondary ionization mass spectrometry to profile aldehydes and ketones [24]. Steroids and sterols in umbilical cord blood have been analyzed using Girard reagents [25] and the conversion of progesterone to 16a-hydroxypregn-4-ene-3,20-dione and androst-4-ene-3,17-dione has been studied in human placenta using Girard reagent [26]. This body of work highlights the utility of Girard reagents in the analysis of molecules containing carbonyl moieties. To our knowledge, this paper is the first account describing the use of Girard reagents and electrospray ionization mass spectrometry to analyze oxidized lipids from human placenta.
Reaction of aldehydes with Girard reagent T provided the added advantage that the product of the reaction contains an extremely weak carbon-nitrogen bond. Thus, these Girard reagent T conjugates predictably fragment with a loss of trimethylamine under mass spectrometric collision induced dissociation conditions (Figure 1).
This predictable fragmentation in the mass spectrometer allows samples to be profiled for all species that react with the derivatization reagent. Molecules that react with Girard reagent T will fragment losing 59 Da (the trimethylamine moiety). This equates to a ‘neutral’ loss of 59 Da for the aldehyde-derivative. Thus, with biological samples it is possible to ‘profile’ a diseased sample (e.g., hypertensive placentas) versus a control sample (e.g., normal placenta) to explore for unexpected oxidation products. This profiling has also been used recently by Johnson and co-workers in their studies of ketosteroids [19].
Thus, the ultimate aims of this work were to develop and qualify a method to quantitate seven key oxidized lipids in placenta and to develop a secondary method to profile placental samples for other oxidized lipids.
2. Materials and Methods
2.1 Reagents
Heptanal was purchased from Alfa Aesar (Ward Hill, MA). Hexanal, Girard Reagent T (GirT), and Heptafluorobutyric Acid were purchased from Acros Organics/Fisher (Hampton, NH). The 4-hydroxyhexenal and 4-hydroxynonenal were purchased from Cayman Chemical Company (Ann Arbor, MI). Acrolein was purchased from MP Biomedicals (Irvine, CA). Nonanal and the Malondialdehyde tetrabutylammonium salt was purchased from Fluka Chemical Company (Switzerland). All other reagents were purchased and used as received from Fisher Scientific (Hampton, NH).
2.2 Placental Samples
In the Department of Pathology at University of Texas Southwestern Medical Center, approximately 100 mg (each) of chorionic tissue was collected from de-identified freshly delivered placentas. Following procurement, the tissue samples were immediately frozen on dry ice and stored at -80 ± 10 ºC until further processing. IRB approval was obtained for this study and patients signed informed consent to be part of the study.
2.3 Solution Standards
Solutions of the lipid aldehydes were prepared in 100% acetonitrile at a concentration of ~ 1000 ng/mL. These stock solutions were used for the direct infusion experiments to determine the observed mass-to-charge ratios under electrospray ionization conditions. This stock solution was also used as a chromatographic standard during the LC-MS/MS method development. Serial dilutions of the 1000 ng/mL stock standard in acetonitrile produced standards with concentrations of 10 ng/mL to 1000 ng/mL. These lipid aldehyde standards were reacted with Girard reagent T and worked up to produce calibration and quality control standards. All lipid aldehyde solutions were stored at -10 to -25 °C when not in use.
2.4 Placental Sample Preparation
Placental samples were thawed and ground in an aqueous environment (100 mg placenta in 1.0 mL Milli-Q water). Following grinding, the suspended tissue was centrifuged (12,000 rpm for 10 min at room temperature) and the supernatant was removed and extracted two times with 1 mL of ethyl acetate. The ethyl acetate layers were combined and the solvent was removed using a vacuum centrifuge. The residue was dissolved in 100 µL of acetonitrile prior to derivatization.
2.5 Derivatization Reactions
Derivatization reactions were conducted by incubation of the solutions of lipid aldehydes or placenta (100 mL) with 12 mM GirT (100 mL). GirT (~50mg) was dissolved using 20 mL of acetonitrile and then acidified with 400 µL of formic acid. The reactions were allowed to proceed for at least 2 hours at room temperature and then put under a steady stream of N2 to complete dryness; if not all samples were under N2 they were put in the freezer (-10 to -25 °C) in order to halt the reaction. The samples were then reconstituted with 300 mL of water:methanol (98:2, v/v) and mixed well. The standards were then analyzed by LC-MS/MS with positive electrospray ionization and a 20 mL injection volume.
2.6 Direct-Infusion Experiments
The ions associated with each aldehyde were obtained by direct-infusion mass spectrometry on a Sciex API-3000 Tandem Triple-Quadrupole Mass Spectrometer equipped with an electrospray ionization source and interfaced with a Harvard Apparatus syringe pump. The product ions for each of the precursor ions were subsequently determined of the aldehydes.
2.7 HPLC-MS/MS
Chromatographic separation was achieved using a Varian ODS-3 (5m) reversed-phase column (2.1 x 50 mm). The column provided a chromatographic separation of the lipid aldehydes present in the experiments. No guard column was used given the relatively extensive liquid-liquid sample preparation utilized prior to derivatization. Gradient elution in the presence of the volatile ion-pairing reagent heptafluorobutyric acid (HFBA) was found to provide optimal chromatographic peaks and resolution. The HPLC employed was a Shimadzu SIL-HTc module with mobile phase A (MP A) 0.1% HFBA in acetic acid (20 mM), and mobile phase B (MP B) 100% methanol at a flow rate of 0.200 mL/min. The gradient elution program was as follows: 2% MP B for the first 2 minutes; from 2% to 98% MP B in 3 minutes, linear gradient, and hold for 2 minutes; from 98% to 2% MP B in 0.5 minutes, linear gradient, and hold at 2% MP B for 2.5 minutes.
The API-3000 mass spectrometer was used in multiple reaction monitoring (MRM) mode analyzing for the ions determined in the direct infusion experiments. The optimized mass spectral conditions for quantitation included a nebulizer gas setting of 7.0, a curtain gas setting of 12.0 a CAD gas setting of 7.0, an ion spray voltage of 5500 V, and a source temperature of 500 °C. The optimized compound specific mass spectrometer settings are presented in Table 1.
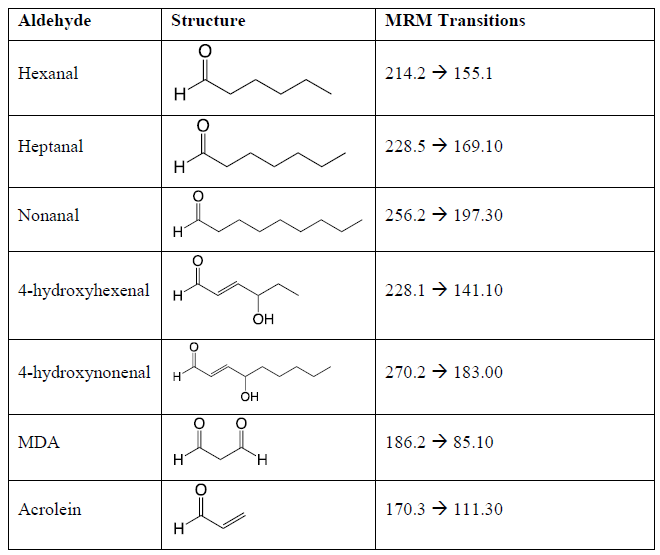
Table 1: The target lipid aldehydes, their structures, the precursor-to-product mass-to-charge ratios for the Girard reagent T derivatized lipid aldehydes, and the optimized compound specific mass spectrometric conditions. Declustering potential = 40V, Entrance potential = 10V, Collision Energy = 30V, and Collision Cell Exit Potential = 10V.
The concentration-response (linearity) of the calibration curve was evaluated by: (1) comparison of each determined calibrator concentration to the concentration calculated by linear regression and (2) examination of the r value. The nominal concentrations of the calibrators were 10, 25, 50, 100, 500 and 1000 ng/mL. Precision and accuracy measurements were determined by analyzing quality control samples at known concentrations of the lipid aldehydes in an aqueous environment. At least three sets of control samples for all concentrations were evaluated.
3. Results
3.1 Direct Infusion Experiments (Optimization of the Mass Spectrometric Conditions
The ions and MS/MS CID fragment patterns associated with the lipid aldehydes were obtained by direct-infusion MS analysis. A unique MS/MS precursor-to-product transition was obtained for each of the derivatized lipid aldehydes hexanal, heptanal, nonanal, 4-hydroxyhexenal, 4-hydroxynonenal, MDA and acrolein (Table 1). It is noteworthy that the MS/MS fragment related to Precursor-59 was observed in all MS/MS CID fragmentation; however, it was not always the most abundant fragment. The most intense MS/MS CID fragment was chosen for targeted quantitation.
3.2 Optimization of the Placental Sample Preparation
A number of different homogenization and extraction methodologies can be found in the analysis of oxidized lipids in biofluids and tissues [27,28]. Four different homogenization - extraction methods were compared to determine the best method for the analysis of oxidized lipids in placental tissue. The following four different sample preparation techniques were investigated:
- Homogenization in water – Chloroform: Methanol Extraction
- Homogenization in 50 mM LiCl aq. – Chloroform:Methanol Extraction
- Homogenization in Chloroform:Methanol – Extraction following addition of water
- Homogenization in water – Ethyl Acetate Extraction
Homogenization in LiCl aq. with Chloroform:Methanol Extraction (Bligh-Dyer method) [29] was consistently found to provide the most efficient extraction.
3.3 Optimization of the Derivatization Reaction
GirT was selected as the derivatization agent because it produces positively charged derivatives and its potential to be used to profile the molecules that contain an aldehyde or ketone. The derivatization procedure was based on previous examples of hydrazone formation with Girard reagents [12]. Acetonitrile was selected as the solvent for the derivatization due to its compatibility with GirT and the oxidized lipids. A small portion of formic acid was used in the reaction to catalyze the formation of the hydrazone and due to its compatibility with mass spectrometry.
The length and the duration of the reaction were examined. Reaction times of 0.5, 1.0, 2.0 and 3.0 hours were investigated using concentrations of the oxidized lipid aldehydes from 10 to 1000 ng/mL, and 12 mM concentration of GirT. The reaction was found to not be complete at 0.5 hours. Reactions of 1, 2 and 3 hours provided similar results to each other, thus a reaction time of 2 hours was selected. The optimal temperature of the reaction was examined. Reaction temperatures of 23ºC (room temperature) and 37ºC were investigated found to produce similar results; therefore, subsequent reactions were conducted at room temperature.
The addition of an antioxidant was investigated to prevent the spontaneous oxidation of lipids during the sample preparation procedure. Reaction in the presence of and in the absence of butylated hydroxytoluene (2,6-di-tert-butyl-p-cresol, BHT, 0.01 M in the reaction), a common antioxidant, were investigated. The presence of BHT did not alter the reaction of the solution standards, thus the presence of an antioxidant was not deemed necessary using this methodology. This is similar to previous results obtained with exhaled breath condensate and oxidized lipids [12].
3.4 Optimization of the Chromatography
Optimal chromatographic resolution and peak shape was achieved using a Varian ODS-3 (5m) reversed-phase column (2.1 x 50 mm) and gradient elution in the presence of the volatile ion-pairing reagent HFBA. Representative chromatograms for each of the lipid aldehydes are presented in Figure 2.
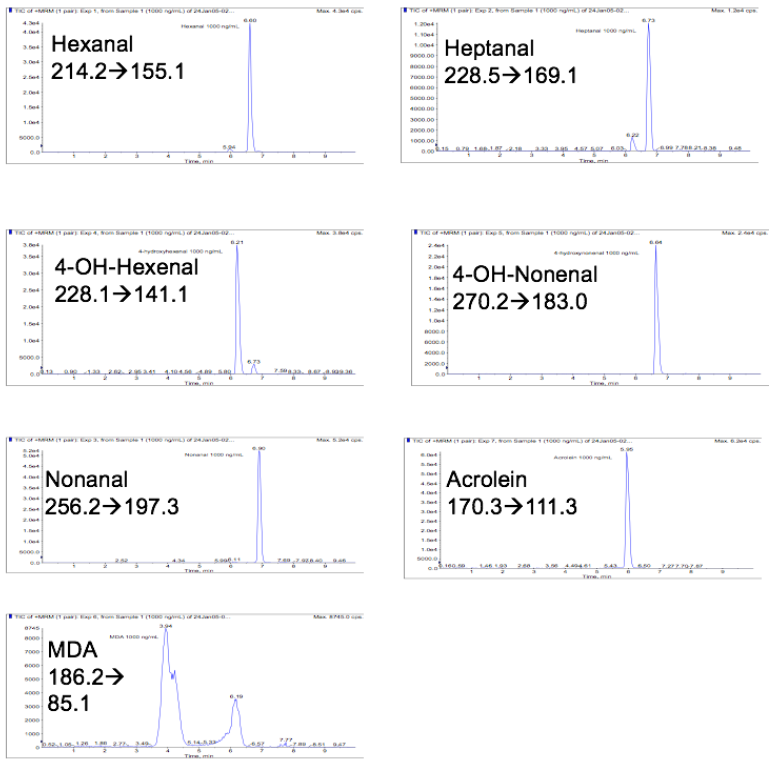
Figure 2: Representative LC-MS/MS chromatograms for Girard reagent T derivatized lipid aldehydes.
3.5 Method Qualification – Linearity
The linear concentration range was determined to be from 10 to 1000 ng/mL for the lipid aldehydes studied. This is the working concentration range for this study. The full range may be greater. A weighted linear regression of (1/×) was used. Interday values for slope, intercept and correlation coefficient values are depicted in Table 2 using data from three separately-prepared calibration curves. It is noteworthy for these initial experiments that an internal standard was not used; however, a spiked lipid aldehyde could be used as an internal standard. The observed accuracy and precision of the method was appropriate for the aims of the study without the use of an internal standard.
Table 2: Method qualification: Concentration-response relationship of Girard reagent T derivatized lipid aldehydes.
|
Analyte |
Slope |
Intercept |
Correlation coefficient (r2) |
|
Hexanal |
221 |
4770 |
0.962 |
|
Heptanal |
81.0 |
125 |
0.968 |
|
Nonanal |
329 |
-561 |
0.956 |
|
4-hydroxyhexenal |
260 |
434 |
0.999 |
|
4-hydroxynonenal |
158 |
4100 |
0.997 |
|
MDA |
229 |
7030 |
0.989 |
|
Acrolein |
451 |
4600 |
0.999 |
3.6 Method Qualification – Accuracy
Intraday accuracy measurements were determined through the analysis of the percent relative error (% RE) of quality control standards at concentrations of 10, 25, 50, 100, 500 and 1000 ng/mL. Three replicates at each concentration was prepared across three days for method qualification (n = 9 per concentration per analyte). The % REs at each calibration point was determined and they are presented in Table 3. The % REs varied between 0.00% and -24.5% for accuracy determinations of concentrations between 25 ng/mL through 1000 ng/mL. It is noteworthy that 4 of 35 accuracy values fell outside of the FDA Guidance for Industry on Bioanalytical Method Validation recommended values of ±15%. The % REs varied between -1.57% and 18.6% at the 10 ng/mL concentration (i.e., the functional lower limit of quantitation for this analysis). It is noteworthy that none of the accuracy values at the lower limit of quantitation fell outside of the FDA Guidance for Industry on Bioanalytical Method Validation (September 2013) recommended accuracy values of ±20% at the lower limit of quantitation. Although there were a minor number of accuracy determinations (4 of 42 in total) that fell outside of the recommended accuracy tolerances of the FDA Guidance for Industry on Bioanalytical Method Validation, the method was determined to be suitable for its intended use because it was developed to determine large changes in specific aldehyde content.
Table 3: Method qualification: Concentration-response relationship of Girard reagent T derivatized lipid aldehydes.
|
Analyte |
10 ng/mL |
25 ng/mL |
50 ng/mL |
100 ng/mL |
500 ng/mL |
1000 ng/mL |
|
Hexanal |
4.03% |
-12.5% (12.7) |
14.3% |
-0.60% |
-8.13% |
3.90% |
|
Heptanal |
-15.8% (27.8) |
-13.2% (7.81) |
13.3% |
18.0% |
-0.87% (14.7) |
-1.43% (3.59) |
|
Nonanal |
-3.67% (16.4) |
4.53% |
0.00% |
3.43% |
-8.00% (17.1) |
3.43% |
|
4-hydroxyhexenal |
-14.7% (16.2) |
-14.8% (5.89) |
-3.6% |
13.0% |
2.9% |
-2.1% |
|
4-hydroxynonenal |
-1.57% (15.4) |
8.27% |
22.5% |
12.1% |
2.33% |
-3.73% (4.13) |
|
MDA |
18.6% |
-24.5% (21.3) |
7.40% |
18.3% |
2.27% |
-2.80% (16.1) |
|
Acrolein |
3.70% |
-1.33% (4.63) |
-5.00% (16.0) |
-2.50% (11.1) |
2.13% |
-0.57% (1.09) |
3.7 Method Qualification – Precision
Intraday precision measurements were determined by analyzing QC samples of known concentrations of the lipid aldehydes. Three replicates at each concentration was prepared across three days for method qualification (n = 9 per concentration per analyte). The percent relative standard deviations (% RSDs) at each quality control level is presented in Table 3. The % RSDs varied between 1.09% and 21.3% for precision determinations of concentrations between 25 ng/mL through 1000 ng/mL. It is noteworthy that 6 of 35 of the precision values fell outside of the FDA Guidance for Industry on Bioanalytical Method Validation (September 2013) recommended values of ±15%. The % RSDs varied between 14.1% and 26.9% for the 10 ng/mL concentration (i.e., the functional lower limit of quantitation for this analysis. It is noteworthy that 2 of 7 of the precision values at the lowe limit of quantitation fell outside of the FDA Guidance for Industry on Bioanalytical Method Validation (September 2013) recommended values of ±20% at the lower limit of quantitation. Although there were a minor number of accuracy determinations (8 of 42 in total) that fell outside of the recommended precision tolerances of the FDA Guidance for Industry on Bioanalytical Method Validation, the method was determined to be suitable for its intended use because it was developed to determine large changes in specific aldehyde content.
3.8 Method Qualification – Limit of Detection
The limit of detection was not rigorously examined. All seven analytes demonstrated a signal-to-noise ratio significantly greater than three-to-one at the 10 ng/mL level. Thus, the actual limit of detection may be much lower than 10 ng/mL (in solution).
3.9 Quantitation of Oxidized Lipids in Placental Tissue Samples
A pilot set of placental samples was analyzed for content of the seven key oxidized lipid components using the LC-MS/MS methodology. The pilot set of placental samples included 2 from normotensive mothers and 20 from mothers with hypertension during pregnancy. The quantitation was conducted using an external solution calibration curve because this application of the method was designed to directly compare the two sets of samples. Table 4 depicts the results of this pilot set of placental samples. These values are referenced to the initial weight of placental tissue used.
Table 4: Quantitation of oxidized lipids in control placenta samples and placenta samples from women with preeclampsia (PE). Concentrations are expressed as µg of analyte per gram of placenta processed.
|
Sample Type |
n |
Hexanal (µg/g) |
Heptanal (µg/g) |
Nonanal (µg/g) |
4-hydroxyhexenal (µg/g) |
4-hydroxynonenal (µg/g) |
MDA (µg/g) |
|
Control |
2 |
4.99 |
0.0525 |
0.150 |
0.153 |
5.10 |
13.43 |
|
PE samples |
20 |
3.07 |
0.0243 |
0.157 |
0.141 |
3.31 |
8.04 |
3.10 Profiling of Placental Tissue Samples for Oxidized Lipid Content (Untargeted Analyses)
The placental tissue samples were then analyzed employing the neutral loss scan (Precursor-59). The metabolite ID software package (Applied Biosystems, Foster City, CA) was then used to profile the differences in the placental samples from normotensive and hypertensive mothers. The samples were compared on a one-by-one basis over the retention time window 2-8 minutes. Differences were scanned over the mass-to-charge ratios 80 – 400. Seven nominal masses were consistently observed: m/z of 299, 236, 205, 214, 200 158, and 144. Investigations continue on the molecular identities of these species. Interestingly, the m/z of 200 corresponds to the molecular weight of pentanal, a lipid oxidation product that has not been previously implicated in oxidative stress disorders.
4. Discussion
A new method has been developed and qualified to analyze seven key oxidized lipids in placental tissue. The method employs derivatization of the lipid aldehydes with GirT to increase the sensitivity. The method was found to be sufficiently accurate, precise, and sensitive for quantitation. The use of the derivatization agent GirT was found to substantially increase the number of lipid aldehyde species that can be readily quantitated in biofluids using electrospray ionization and LC-MS/MS.
An extension of this method using neutral loss scanning allows profiling of placental samples for unexpected oxidized lipids (untargeted lipid aldehyde-omics). This methodology provides a unique top-down approach to profile biological samples for oxidized lipids. The neutral-loss scanning technique is applicable to any analyte that reacts with GirT, thus this methodology provides a technique that should be applicable to profiling any biofluid or tissue for unexpected products of lipid oxidation.
This method was used to analyze a small number of placental samples from normotensive and hypertensive mothers. It was determined that placental tissue from mothers with preeclampsia had relatively lower amounts of oxidized lipids when compared to control placental tissue. This differential expression may provide insight into potential biomarkers or biological profile of lipid aldehyde expression in preeclampsia.
Author Contributions:
Conceptualization, W.C.P and D.R.; methodology, W.C.P. and S.M.S.; formal analysis, S.M.S.; writing—original draft preparation, W.C.P and D.R..; writing—review and editing, W.C.P and D.R. All authors have read and agreed to the published version of the manuscript.
Funding:
This research was funded by the Jerry H. Hodge School of Pharmacy at Texas Tech University health Sciences Center.
Conflicts of Interest:
The authors declare no conflict of interest.
References
- Allen RE, Rogozinska E, Cleverly K, et al. Abnormal blood biomarkers in early pregnancy are associated with preeclampsia: a meta-analysis. European Journal of Obstetrics & Gynecology and Reproductive Biology 182 (2014): 194-201.
- Ferguson KK, Meeker JD, McElrath TF, et al. Repeated measures of inflammation and oxidative stress biomarkers in preeclamptic and normotensive pregnancies. American Journal of Obstetrics and Gynecology 216 (2017): 527-e1.
- Petla LT, Chikkala R, Ratnakar KS, et al. Biomarkers for the management of pre-eclampsia in pregnant women. The Indian Journal of Medical Research 138 (2013): 60.
- Masoura S, Kalogiannidis IA, Gitas G, et al. Biomarkers in pre-eclampsia: a novel approach to early detection of the disease. Journal of Obstetrics and Gynaecology 32 (2012): 609-616.
- Cuffe JS, Holland O, Salomon C, et al. Placental derived biomarkers of pregnancy disorders. Placenta 54 (2017): 104-110.
- Roberts JM, Hubel CA. Is oxidative stress the link in the two-stage model of pre-eclampsia?. The Lancet 9181 (1999): 788-789.
- Poranena AK, Ekblad U, Uotila P, et al. Lipid peroxidation and antioxidants in normal and pre-eclamptic pregnancies. Placenta 17 (1996): 401-405.
- Mutlu-Türkoglu Ü, Ademoglu E, Ibrahimoglu L, et al. Imbalance between lipid peroxidation and antioxidant status in preeclampsia. Gynecologic and Obstetric Investigation 46 (1998): 37-40.
- Esterbauer H, Schaur RJ, Zollner H. Chemistry and biochemistry of 4-hydroxynonenal, malonaldehyde and related aldehydes. Free radical Biology and Medicine 11 (1991): 81-128.
- Liu J, Yeo HC, Doniger SJ, et al. Assay of aldehydes from lipid peroxidation: gas chromatography–mass spectrometry compared to thiobarbituric acid. Analytical Biochemistry 245 (1997): 161-166.
- Scott MD, Eaton JW. Free radical toxicology. ed. by Wallace KB, 401–420.
- Andreoli R, Manini P, Corradi M, et al. Determination of patterns of biologically relevant aldehydes in exhaled breath condensate of healthy subjects by liquid chromatography/atmospheric chemical ionization tandem mass spectrometry. Rapid Communications in Mass Spectrometry 17 (2003): 637-645.
- Stalikas CD, Konidari CN. Analysis of malondialdehyde in biological matrices by capillary gas chromatography with electron-capture detection and mass spectrometry. Analytical Biochemistry 290 (2001): 108-115.
- Deighton N, Magill WJ, Bremner DH, et al. Malondialdehyde and 4-hydroxy-2-nonenal in plant tissue cultures: LC-MS determination of 2, 4-dinitrophenylhydrazone derivatives. Free Radical Research 27 (1997): 255-265.
- Orhan H, van Holland B, Krab B, et al. Evaluation of a multi-parameter biomarker set for oxidative damage in man: increased urinary excretion of lipid, protein and DNA oxidation products after one hour of exercise. Free Radical Research 38 (2004): 1269-1279.
- Wu T, Rifai N, Roberts LJ, et al. Stability of measurements of biomarkers of oxidative stress in blood over 36 hours. Cancer Epidemiology and Prevention Biomarkers 13 (2004): 1399-1402.
- Stenlake JB, Williams WD. The Spectrophotometric Determination Of Αβ-Unsaturated Aldehydes And Ketones With Girard-T Reagent: Part I. EssentialOils. Journal of Pharmacy and Pharmacology 9 (1957): 900-907.
- Stenlake JB, Williams WD. The Spectrophotometric Determination Of Αβ-Unsaturated Aldehydes And Ketones With Girard-T Reagent: Part II. Ketosteroids. Journal of Pharmacy and Pharmacology 9 (1957): 908-913.
- Johnson DW. Ketosteroid profiling using Girard T derivatives and electrospray ionization tandem mass spectrometry: direct plasma analysis of androstenedione, 17-hydroxyprogesterone and cortisol. Rapid Communications in Mass Spectrometry: An International Journal Devoted to the Rapid Dissemination of Up-to-the-Minute Research in Mass Spectrometry 19 (2005): 193-200.
- Griffiths WJ, Liu S, Alvelius G, et al. Derivatisation for the characterisation of neutral oxosteroids by electrospray and matrix-assisted laser desorption/ionisation tandem mass spectrometry: the Girard P derivative. Rapid Communications in Mass Spectrometry 17 (2003): 924-935.
- Griffiths WJ, Alvelius G, Liu S, et al. High-energy collision-induced dissociation of oxosteroids derivatised to Girard hydrazones. European Journal of Mass Spectrometry 10 (2004): 63-88.
- Gouw JW, Burgers PC, Trikoupis MA, et al. Derivatization of small oligosaccharides prior to analysis by matrix-assisted laser desorption/ionization using glycidyltrimethylammonium chloride and Girard's reagent T. Rapid Communications in Mass Spectrometry 16 (2002): 905-912.
- Lai CC, Tsai CH, Tsai FJ, et al. Rapid monitoring assay of congenital adrenal hyperplasia with microbore high-performance liquid chromatography/electrospray ionization tandem mass spectrometry from dried blood spots. Rapid Communications in Mass Spectrometry 15 (2001): 2145-2151.
- Ross MM, Kidwell DA, Colton RJ. Selective detection of aldehydes and ketones by derivatization/secondary ion mass spectrometry. International Journal of Mass Spectrometry and Ion Processes 63 (1985): 141-148.
- Eberlein WR. Steroids and sterols in umbilical cord blood. The Journal of Clinical Endocrinology & Metabolism 25 (1965): 1101-1118.
- Little B, Shaw A, Purdy R. The conversion of progesterone to 16α-hydroxy-pregn-4-ene-3, 20-dione and androst-4-ene-3, 17-dione by human placenta in vitro. European Journal of Endocrinology 43 (1963): 510-520.
- Jain S, Jayasimhulu K, Clark JF. Metabolomic analysis of molecular species of phospholipids from normotensive and preeclamptic human placenta electrospray ionization mass spectrometry. Front Biosci 9 (2004): 3167-3175.
- Han X, Yang J, Cheng H, et al. Toward fingerprinting cellular lipidomes directly from biological samples by two-dimensional electrospray ionization mass spectrometry. Analytical Biochemistry 330 (2004): 317-331.
- Bligh EG, Dyer WJ. A rapid method of total lipid extraction and purification. Canadian Journal of Biochemistry and Physiology 37 (1959): 911-917.
