Clinical and Laboratory-based Diagnosis in Cases Suspected with COVID-19; An Updated and Comprehensive Systematic Review Study
Article Information
Nazila Bahmaie1a†, Mehrnaz Ajorloo1b†, Maryam Mohammadbeigi2a‡, Parisa Abedi Elkhichi2b‡, Sheida Alizadeh3a‡, Saeed Soroush3b‡, Fatemeh Rajabi3c‡, Elham Nouri3d‡, Pourandokht Farhangian3e‡, Nasim Mohammadi3f‡, Alireza Mohammadyari 3g‡, Maryam Mohammadi3h‡, Mohammad Javad Hajkazemi3i‡, Masoud Shamohammadi3j‡, Mahnoush Bahrampour3k‡, Samin Rahimi3l‡, Mahsa Jalilinezhad3m‡, Sahar Serajian3n‡, Zahra Dorosti3o‡, Samira Lorestani3p‡, Seyed Mohammad Mohyeddin Kazemeini3q‡, Sina Ekhlasi3r‡, Sheida Janati3s‡, Nima Rezaei4a*
1a Department of Allergy and Immunology, Faculty of Medicine, Graduate School of Health Sciences, Near East University (NEU), Nicosia, Northern Cyprus, Cyprus
1b Faculty of Medicine, Shiraz University of Medical Sciences (SUMS), Shiraz, Iran
2a Department of Microbiology and Immunology, Faculty of Medicine, Qazvin University of Medical Sciences (QUMS), Qazvin, Iran
2b Medical Microbiology Research Center, Qazvin University of Medical Sciences (QUMS), Qazvin, Iran.
3a Department of Medical Microbiology and Virology, Faculty of Medicine, Shiraz University of Medical Sciences (SUMS), Shiraz, Iran
3b Faculty of Medicine, Guilan University of Medical Sciences (GUMS), Guilan, Iran
3c Department of Medical Genetics, Faculty of Medical Sciences, Tarbiat Modares University, Tehran, Iran
3d Clinical Diagnosis Laboratory, Shahid Beheshti University affiliated Hospital, Zanjan University of Medical Sciences, Zanjan, Iran
3e Department of Medical Genetics and Molecular Medicine, Faculty of Medicine, Zanjan University of Medical Sciences (ZUMS), Zanjan, Iran
3f Faculty of Pharmacology, Kerman University of Medical Sciences (KUMS), Kerman, Iran
3g Department of Biology, Faculty of Basic Sciences, Hamadan Branch, Islamic Azad University, Hamadan, Iran
3h Department of Microbiology, Faculty of Basic Medical Sciences, Tehran Medical Sciences Branch, Islamic Azad University, Tehran, Iran
3i Faculty of Medicine, Zanjan University of Medical Sciences (ZUMS), Zanjan, Iran
3j Department of Basic Sciences, Faculty of Veterinary Medicine, Razi University of Kermanshah, Kermanshah, Iran
3k Faculty of Pharmacology Sciences, Tehran Medical Sciences Branch, Islamic Azad University, Tehran, Iran
3l Department of Biology, Faculty of Basic Sciences, University of Maragheh, Maragheh, East Azerbaijan, Iran
3m Department of Stem Cells and Developmental Biology, Cell Science Research Center, Royan Institute for Stem Cell Biology and Technology, ACECR, Tehran, Iran
3n Department of Molecular System Biology, Cell Science Research Center, Royan Institute for Stem Cell Biology and Technology, ACECR, Tehran, Iran.
3o Department of Medical Parasitology, School of Medicine, Isfahan University of Medical Sciences, Isfahan, Iran
3pDepartment of Medical Parasitology, Faculty of Medicine, Zanjan University of Medical Sciences (ZUMS), Zanjan, Iran
3q Department of Biology, Faculty of Sciences, Science and Research Branch, Islamic Azad University, Tehran, Iran
3r Department of Microbiology, Faculty of Basic Sciences, Tehran North Branch, Islamic Azad University, Tehran, Iran
3s Immunology Research Center, Institute of Immunology and Infectious Diseases, ACECR, Tehran, Iran
4a*Research Center for Immunodeficiencies, Children’s Medical Center, Tehran University of Medical Sciences (TUMS), Tehran, Iran
† These authors contributed equally to this work.
‡ These authors contributed equally to this work.
*Corresponding author: Nima Rezaei, Research Center for Immunodeficiencies, Children’s Medical Center, Tehran University of Medical Sciences (TUMS), Tehran, Iran
Received: 08 February 2022; Accepted: 15 February 2022; Published: 17 March 2022
Citation: Nazila Bahmaie, Mehrnaz Ajorloo, Maryam Mohammadbeigi, Parisa Abedi Elkhichi, Sheida Alizadeh, Saeed Soroush, Fatemeh Rajabi, Elham Nouri, Pourandokht Farhangian, Nasim Mohammadi, Alireza Mohammadyari, Maryam Mohammadi, Mohammad Javad Hajkazemi, Masoud Shamohammadi, Mahnoush Bahrampour, Samin Rahimi, Mahsa Jalilinezhad, Sahar Serajian, Zahra Dorosti, Samira Lorestani, Seyed Mohammad Mohyeddin Kazemeini, Sina Ekhlasi, Sheida Janati, Nima Rezaei. Clinical and Laboratory-based Diagnosis in Cases Suspected with COVID-19; An Updated and Comprehensive Systematic Review Study. Fortune Journal of Health Sciences 5 (2022): 126-158.
Share at FacebookAbstract
Objectives A novel coronavirus disease 2019 (COVID-19) was identified in Wuhan, China, which quickly involved majority of the countries all around the world. Due to the high rate of mortality and morbidity, needless to say the importance of accurate and early diagnosis, especially in suspected and asymptomatic cases. Hence, in this article, authors tried to provide practical and standardized diagnostic approaches for cases suspected with COVID-19 infection. Material and Methods Data of this review study were collected from 7 search engine/databases, commencing from December 2019 to June 2021 by using 6 keywords according to Medical Subject Headings (MeSH) terms and our inclusion/exclusion criteria. Result Due to various clinical manifestations of COVID-19, and high potential for mutagenicity, identification of suspected patients is of great importance for effective control of infection, and improvement of clinical decisions. Therefore, medical history of the patients, clinical signs and symptoms, chest computational tomography, serological and molecular diagnosis can be effective in faster identification of mentioned patients. In spite of the fact that molecular tests have been considered as the gold standard for diagnosis of COVID-19, but there is still high rate of false-negativity. Then, combinative usage of the complementary tests can reduce any misinterpretations for suspected cases. Conclusion Screening for suspected cases in the shortest possible turnaround time is dependent on the appropriate diagnostic approaches. Subsequently, this allows physicians immediately provide proper medical interventions for suspected patients who are at greater risk for developing more serious complications than COVID-19 like severe nosocomial infections.
Keywords
Coronaviruses, COVID-19, Diagnosis, Detection Rate, NAAT, Suspected patients
Coronaviruses articles, COVID-19 articles, Diagnosis articles, Detection Rate articles, NAAT articles, Suspected patients articles
Article Details
1. Introduction
From December 2019, a burgeoning rate of population have been involved in a highly contagious type of a severe respiratory disorder (Coronavirus 2019) in the Huanan seafood wholesale market of Wuhan, Hubei province, China [1, 2]. On 11th of February 2020, it was renamed as Severe Acute Respiratory Syndrome related Coronavirus-2 (SARS-CoV-2) by International Committee on Taxonomy of Viruses (ICTV) [3-5]. Phylogenetic-based studies proved common features among SARS-CoV-2, Severe Acute Respiratory Syndrome-causing-Corona Virus (SARS-CoV), and Middle East Respiratory Syndrome-causing-Corona Virus (MERS-CoV) [6-8]. Socioeconomical effects of SARS-CoV-2 outbreak made World Health Organization (WHO) to announce a rapidly-spreading pandemic from 11th of March 2020 [7-13], and a societal concern for health managers [14]. No hesitate that an integrative collaboration of scientists is necessary for effective management of this situation by the usage of interdisciplinary frameworks [4, 9, 13, 15-17].
Accordingly, Coronavirus Disease 2019 (COVID-19) in asymptomatic carriers and suspected ones should be immediately diagnosed [14, 18] due to the possibility for rapid developing serious co-nosocomial infections, and misdiagnosis [7, 9, 19-21]. WHO defines “cases of COVID-19” as “a person with the laboratory confirmation of SARS-CoV-2 infection irrespective of clinical signs/symptoms” [4, 9, 22]. Trial seventh version of Diagnosis and Treatment Guidelines for COVID-19 issued by the National Health Committee of the People’s Republic of China declares “suspected” cases with one (of four) items of epidemiological history, or two (of three) items of clinical manifestations, or three items of clinical manifestations with no item of epidemiological history [10, 23], or an acute respiratory tract SARS-CoV-2 infection for less than 14 days, or clinical illness compatible with COVID-19, or asymptomatic carriers in a close contact to the confirmed cases of COVID-19 [24, 25].
Speaking of “suspected cases”, pneumonic people with false results like patients with underlying severe cardiovascular sequelae [26], acute renal disorders, and dead people without ascertained SARS-CoV-2 infection [27-29], pediatrics (as potential carriers of COVID-19 with their imperfect immune system) [30], persons with multi-system inflammation [31, 32], Acute Coronary Syndrome (ACS) [33], and pregnant women undergoing delivery during hospitalization are of great prominence for rapid and accurate diagnosis, too. They are considered as the most problematic challenges for COVID-19, who can spread more viral respiratory co-infections than confirmed cases [7, 11, 16, 34-42].
Diverse clinical signs/symptoms, and similar clinical manifestations with other types of respiratory pathogens (Mycoplasma pneumonia, Chlamydia pneumonia, Adenoviruses, Rhinoviruses, Metapneumonovirus, HKU1, NL63, Influenza type A/B, SARS, and MERS) [4, 7, 43] add erroneous results to diagnostic procedures, and adverse clinical outcomes.
Hence, highly-sensitive and time-preserving diagnostic methods can identify potentially infectious people [44], quantify exact viral load, decrease false-negative results from semi-sensitive tests or from suspected individuals [45-47], reduce the Turnaround Time (TAT) for identification of suspected individuals, immediately provide optimal medical interventions, potentially lower in-hospital spreading of SARS-CoV-2 infection, shorten the length of isolation/medical surveillance for suspected ones, lessen the socioeconomic fear for individuals who were in close contact with them, and help for determination of risk stratification [43, 45, 48-52].
Despite meticulous efforts for the sanitary recommendations, poor level of hygiene in undeveloped countries, and complying with quarantine regulations impose serious socio-medical challenges and extortionate expenditures until complete recuperation. Therefore, accurate and rapid diagnostic strategies can eliminate uncontrolled release of SARS-CoV-2 infection by asymptomatic carriers, and suspected persons [53-55].
Diagnosis of SARS-CoV-2 infection are mainly done by chest Computed Tomography (CT) images, and serological/molecular-based tests [56]. Molecular approaches reveal high analytical accuracy for initial quantitative diagnosis of SARS-CoV-2 infection [49] (like Nucleic Acid Amplification Tests (NAATs) strategies) [57]. Serological approaches (like Enzyme Linked Immunosorbent Assay (ELISA)) are highly dependent on the detection of neutralizing antibodies [9, 54], or viral antigenic proteins after viral exposure [9, 53, 54].
In this systematic literature review study, diagnostic efficacy of those methods for suspected cases with SARS-CoV-2 infection have been analyzed aimed at providing a successful adjustment of laboratory-based data with optimal clinical outcomes, and hopefully increasing life-expectancy for suspected cases with COVID-19.
2. Methodology
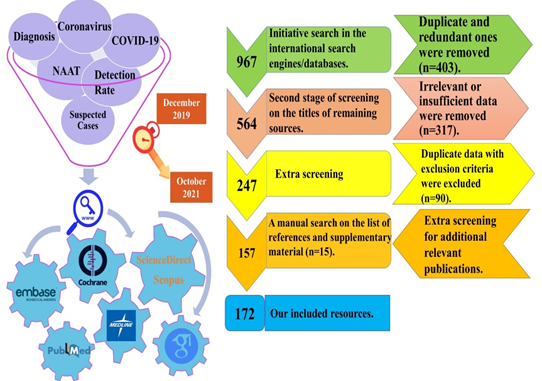
This comprehensive systematic review (systematic literature review) study was performed according to Preferred Reporting Items for Systematic review and Meta-Analysis (PRISMA) statement guideline (https://www.equator-network.org/reporting-guidelines/prisma/) on June 2021 (figure-1).
2.1 Literature Search Strategy and Screening Process
An electronic comprehensive literature review was conducted with time interval commencing from December 2019 to October 2021, by using six main keywords (Figure 1), and four complementary ones (Clinical Diagnosis, AND Laboratory Diagnosis, AND Molecular Diagnosis, AND Serological Diagnosis), based on our inclusion and exclusion criteria.
In order to find potentially-eligible resources, one author independently conducted screening process in three main and one non-electronic backward steps (on the references/supplementary/bibliographies of included articles). Any uncommon points or disagreements were referred to the corresponding author (figure-1).
2.2 Inclusion and Exclusion Criteria
All of the published (original (experimental/non-experimental), review (mini-review/best evidence/narrative review/systematic reviews (and meta-analysis), comparative/cross sectional/cohort/retrospective/observational/commentary/letter to the editors/editorial/opinion/short (rapid) communication/Randomized Clinical Trials (RCTs)/case reports/series) articles were considered in the formats of full-text/full-length, abstract, section of book, chapter, and conference papers in English language. Studies involving suspected human subjects aimed at laboratory-based assessment of SARS-CoV-2 infection through usage of biological samples (whole blood, serum, plasma, Nasopharyngeal (NP)/Oropharyngeal (OP) swabs, Cerebrospinal Fluid (CSF), Bronchoalveolar Lavage (BAL), tracheal secretions, sputum, and saliva), CT, and High Resolution CT (HRCT) were totally included.
In order to report the impact of the most efficient diagnostic method, studies involving human cases suspected with SARS-CoV-2 infection who had previously used any adjuvants, vaccines, anti-viral/herbal/self-therapeutic regimens all were excluded. Studies with irrelevant/insufficient/ambiguous data, lack of data, undefined diagnostic values/diagnostic methods all were excluded.
2.3 Data Extraction
Five independent authors majorly performed data extraction and made forms to collect study characteristics (author name, publication date, study design, used samples, types/subtypes of diagnosis). In case of overlapping data or several published reports from same studies, the authors tried their best to present the most complete data.
2.4 Bio-statistical Analysis
None
2.5 Ethical Statement
According to the structure and type of study, there is no need to register for Research Ethical Committee (REC). All of the data supporting the findings of this study, are openly available in the context of this study.
3. Results
3.1 Laboratory Guidelines
Polymerase Chain Reaction (PCR) is one of the most sensitive diagnostic approaches, detecting viral ribonucleic acid of SARS-CoV-2 infection in NP and OP swabs of asymptomatic carriers or suspected cases. Due to false-negative results in PCR, sputum, and saliva are currently-considered specimens with better detection rate in comparison with OP or NP swabs [58]. Because of hurdles in sputum sampling for diagnosis of SARS-CoV-2 infection [47, 59, 60], feces, and blood sampling are also of diagnostic value for increasing detection rate of SARS-CoV-2 infection in suspected persons [61-63].
Therefore, rapid laboratory assessment of appropriate samples from cases suspected with SARS-CoV-2 infection through real-time Reverse Transcriptase-quantitative PCR (rRT-qPCR), and evaluation of other co-respiratory infections will be recommended in order to reduce morbidities/co-morbidities [64], and set up the most practical precautionary or therapeutic approach [65]. Moreover, postmortem sampling through an autopsy should be assessed by PCR for decedent cases with no ascertained SARS-CoV-2 infection. For virological confirmation or exclusion, swab collections of primary bronchi from two categories of suspected persons are evaluated (those with initiative negative NP and OP swabs who represent significant pulmonary findings, and those with initiative positive NP and OP swabs with or without presentation of significant pulmonary findings) [37].
As the number and variety of available tests for detection of SARS-COV-2 infection are increasing, there is an imperative need to more comprehensive understanding on variety of the laboratory-based diagnosis of viral infections, different types of recruited samples, and viral genomic mutations to optimize single-objective diagnostic tests while maintaining the diagnostic sensitivity and specificity. Due to the possibility of repetitive mutations, regular sequencing of the evolved virus can specify structural and functional changes in the primer and probe binding sites, sensibly contributing to decipher complex immunopathogenesis, improving the epidemiological studies, and preventing from more transmission/emergence of recurrent infections.
Moreover, by decreasing the accurate time required for better management of patients with COVID-19, overcoming the challenges associated with the development of rapid Point Of Care (POC) diagnostic should not be underestimated [66, 67]. Due to the ambiguous pathogenesis of newly-mutated Coronaviruses, those two broad diagnostic categories (molecular and serological assessments) will not be interpreted solely [68].
3.2 Molecular Detection of COVID-19
Nucleic acid hybridization, rRT-qPCR, viral genome sequencing, and Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR) have been endorsed as gold standard molecular approaches for the initial diagnosis of SARS-CoV-2 infection [69-71], due to availability, accuracy, and efficiency [53, 72]. Among them, highly-specific multiplex RT-PCR is considered as the most practical test for rapid detection of low concentrations of SARS-CoV-2 RNA [73-78].
There is a co-relation between day of onset of symptoms and viral RNA levels (Cycle threshold (Ct)). As positive results from rRT-qPCR can be still reached 6 to 8 days after the loss of transmissibility, viral viability and transmissibility should not be underestimated for an accurate clinical interpretation. Two consecutively-negative results from RT-PCR separated by at least one day interval, confirm hospital discharge for previously-confirmed patients, and discontinuation of quarantine situation for cases suspected with SARS-CoV-2 infection [79-82].
Targeted genes for molecular detection of SARS-CoV-2 infection include Nucleocapsid (N), Envelope (E), Spike (S), Membrane (M), and RNA dependent RNA polymerase (RdRp) genes [83-85]. For clinically-confirmed patients with COVID-19 and negative results in molecular tests, these tests should be consecutively repeated to be synchronized with clinical manifestations [86, 87]. Results from investigation of cases suspected with SARS-CoV-2 infection and similar symptoms between COVID-19 and other common co-respiratory infections, highlight the necessity of differential diagnosis through RT-PCR, supporting efficient anti-viral therapy and patient care [88-92].
Alteri C et al., [45] conducted viral quantification through a droplet digital PCR (ddPCR)-based assay (targeting RdRp and host RNAse P) on NP swabs of 55 cases suspected with SARS-CoV-2 infection and negative rRT-qPCR results. Among them, 19 persons (34.5%) showed positivity for SARS-CoV-2 infection, and IgG through ddPCR, and chemiluminescent microparticle immunoassay, respectively. Chest CT images in 73.7% of that population showed severe COVID-19 manifestations, introducing ddPCR as a sensitive, and complementary diagnostic approach for cases who have low viral copy number in very early stages of viral replication, and negative results in contextual PCR [45].
He J-L et al., [93] serially conducted RT-PCR tests on 82 individuals clinically-suspected with SARS-CoV-2 infection outside of Wuhan city. They reported 79% sensitivity, and 100% specificity for individuals being initially diagnosed positive with RT-PCR. Additionally, CT images revealed 77% sensitivity for positive patients, and 96% specificity among studied population. They concluded that an initial RT-PCR followed by CT images, can potentially reduce false-negativities in RT-PCR results for cases suspected with SARS-CoV-2 infection in the regions out of epidemiologic center [93].
Zhu W et al., [94] in their study, investigated 116 individuals clinically-suspected with SARS-CoV-2 infection outside of Hubei province (Anhui), and reported 32 clinically-confirmed patients with SARS-CoV-2 infection by chest CT images on admission to Emergency Department (ED). 67%, and 40% of negative cases were coping with fever, and Ground-Glass Opacity (GGO), respectively. They demonstrated that a combination of epidemiological features, laboratory tests, and chest CT findings confirm viral infection in the cases suspected with SARS-CoV-2 infection [94].
In a cross-sectional study conducted by Datta A et al., [95] diagnostic value of HRCT with RT-PCR was evaluated on 114 clinically-suspected cases with SARS-CoV-2 infection in Bangladesh. 91.22% of patients showed GGO with no consolidation in chest CT images. Bilateral chest findings, and vascular thickening were reported for 94%, and 66.66% of the patients, respectively. 96 of 114 patients showed positivity in their RT-PCR results. 90 people out of 96 cases revealed positive chest findings. Among 18 persons with negative results in RT-PCR, 14 people showed positivity in chest CT findings, demonstrating that chest CT could be of high sensitivity for primary diagnosis of SARS-CoV-2 infection when early detection or early clinical decision is prioritized [95].
Kuzan T.Y et al., [96], compared the diagnostic efficacy and accuracy of chest findings to RT-PCR results in first admission to the hospital among routine laboratory-confirmed (69 clinically-suspected cases with SARS-CoV-2 infection and dry cough, fever, bilateral multi-lobe involvement, and patchy shapes), and clinically-diagnosed patients with COVID-19 (51 cases with dyspnea, bronchial wall thickening, and GGO). Due to the high sensitivity and low specificity of chest CT, a combination of clinical features, chest CT images, and laboratory tests are highly recommended to differentiate any similarities between results from laboratory, and clinical investigations [96].
In a recently-conducted retrospective study, positive results of molecular tests were reported in 12 out of 28 cases suspected with SARS-CoV-2 infection. As there were no significant differences for laboratory results, and chest CT results (pure/mixed GGO, bilateral lung involvement, and rounded/patchy/linear opacities) among the studied population, RT-PCR tests are still efficacious for confirmed diagnosis of SARS-CoV-2 infection [97].
In a multi-center retrospective study, Miao C et al., investigated diagnostic criteria for chest CT images, and RT-PCR test (on NP/sputum samples) of cases suspected with SARS-CoV-2 infection [98], reporting 76 cases as negative group, and 54 cases as positive group (or initially-negative and subsequently-positive group (with one day interval)).
Despite similar clinical manifestations and low sensitivity, combination of common features (GGO with bilateral pulmonary distribution (57.4% of positive group), GGO with pleural distribution (55.6% of positive group), GGO with bilateral pulmonary and peripleural distributions (48.1% of positive group), and GGO with bilateral pulmonary distribution, the crazy-paving pattern, and pleural distribution (22.2% of positive group)) led to increased specificity (approximately 99% for GGOs with the crazy-paving pattern and bilateral pulmonary distribution), and reduced overlaps in the clinical manifestations common with viral pneumonia. Hence, repeated sampling for molecular tests, and immediate isolation for successful differential diagnosis of SARS-CoV-2 infection are recommended [98]. Case report/series studies on cases suspected with SARS-Cov-2 infection are collected in table 1.
Table 1: Case studies on cases suspected with SARS-Cov-2 infection
Giannitto C et al., [105] investigated 337 cases suspected with SARS-CoV-2 infection with initially-negative results from their molecular tests on BAL/NP samples to weigh up diagnostic value of pneumonic manifestations acquired from chest CT scans. Accordingly, in a time-interval shorter than five days, patients underwent sampling for first NP/OP swabs, CT, and then second BAL or NP swabs. Multiplex real-time RT-PCR assay for simultaneous detection of N, E, and RdRp target genes showed negativity for 87 cases in their first NP sampling (excluding 19 cases with no second round of sampling). Of 68 main participants, 48 cases showed negativity in their second sampling. Among them, there were 24 cases suspected with pneumonia in analysis of CT images. GGO, and positivity in second round of sampling were reported in more than 50% of lung patterns, and 58% of this population. Sensitivity, specificity, and accuracy of CT images combined with second round of molecular tests, were reported 100%, 79%, and 85%, respectively. Then, combinative interpretation of CT-based approaches with molecular tests, provides differential diagnosis of SARS-CoV-2 infection from other viral or bacterial pneumonia, and early identification of false-negative patients [105].
To sum up, for having a validated molecular detection of SARS-CoV-2 infection, at least one of the following criteria must be met: 1) positive NAATs results for at least two different genomic locations of COVID-19, 2) verification of at least one genomic site for COVID-19, being proved by sequencing. If cases suspected with SARS-CoV-2 infection have consecutively-negative results in two days (with one day interval) from molecular investigations, they can be discharged. In other words, for suspected cases with a history of close contact with clinically-confirmedpatients, molecular detection of SARS-CoV-2 should be considered repeatedly at least two times from appropriate sample [106] in the proper time/anatomical site for reducing any misinterpretations, and qualifying therapeutic approaches for SARS-CoV-2 infection [50, 91, 107-109].
3.3 Serological Detection of COVID-19
Serological tests (authorized by Food and Drug Administration (FDA)) can detect specific Immunoglobulin M (IgM) and IgG antibodies against structural units of SARS-CoV-2 [110]. They are wildly used for verification of molecular tests to raise the sensitivity and accuracy of laboratory-based diagnosis especially for cases suspected with SARS-CoV-2 infection [111, 112].
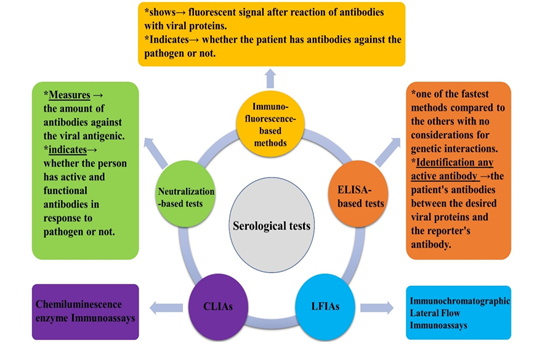
Serological tests identify individuals with therapeutic/prophylactic antibodies, and determine the immune responses to the possible causes of infection [113]. Figure 2 indicates a schematic presentation of serological tests, using plasma, whole blood, or serum specimens for cases suspected with SARS-CoV-2 infection [72, 114-117].
Because of high abundance of N antigenic protein, involvement of S proteins in the attachment to host cell surfaces, and high specificity of the antibodies against Receptor-Binding Domain of S (RBD-S), detection of both antibodies against those antigens results in a highly-sensitive serological diagnosis of SARS-CoV-2 infection. Those serological tests mainly screen the donors of convalescent plasma, specify asymptomatic carriers, determine mortality rate (estimate population exposure rates), demonstrate previous infection, and complementarily verify molecular tests. Therefore, combination of molecular and serological-based tests, may consistently raise the sensitivity for laboratory diagnosis of SARS-CoV-2 infection up to 80–90%, and improve clinical outcomes [81, 118-121]. Simultaneous negativity in IgM and IgG antibodies in cases suspected with SARS-CoV-2 infection, should be followed by repetitive molecular detection or high-throughput viral genome sequencing [122].
H.Zeng et al., [112] investigated the diagnostic value of combinative molecular and serological tests on 71 cases suspected with SARS-CoV-2 infection. They reported increased sensitivity for combined detection (63.38%) compared to solely-measured out molecular tests (46.48%), and serological tests (42.25%), opening promising windows for reduction of false-negativities in cases suspected with SARS-CoV-2 infection [112].
In another retrospective study [123], Jia X et al., investigated 57 cases suspected with SARS-CoV-2 infection. For 24 cases with positive molecular results in primary NP/OP sampling (the first one was confirmed with two additionally-repetitive nucleic acid detection), positive diagnostic rate for combinative antibody detection was 87.50% (more than single-antibody detection). For 33 persons with negative molecular results in primary NP/OP sampling, positive diagnostic rate for combinative antibody detection was 72.73% (more than single antibody detection). Among routine blood tests, only hs-CRP, and Aspartate aminotransferase (AST) showed a statistically-significant difference between studied groups. Regardingly, combinative antibody detection with nucleic acid assessment, and CT images provides an accelerated diagnostic approach with reduced risk of false-negativities among cases suspected with SARS-CoV-2 infection [123].
Negativity in NAAT test, strong epidemiological association, and validated serological assessment in the acute/recovery stages of infection (accompanied by CT-imaging) can support an accurate diagnosis for cases suspected with SARS-CoV-2 infection. Serum samples of asymptomatic patients with the history of close contact with clinically-confirmed patients with SARS-CoV-2 infection, can be stored for sero/epidemiological surveillance [7, 124, 125]. Although interactions with other coronaviruses can be challenging in diagnostic approaches, commercial and non-commercial serological tests are currently underway [82, 126].
The major structurally-specific protein of SARS-CoV-2 reacting with the membrane of host cells is S protein [127], stimulating humoral responses against these proteins which will be measured through western blotting, ELISA, or colorimetric fluorometric outputs (in case of secondary antibody reacting with the bound antibodies). Thus, proper binding of antibody-antigen can justify the folding and spatial shape of the protein, and reducing false-negativities [128]. Moreover, S protein of COVID-19 can be identified by a monoclonal antibody known as CR3022 (for research purposes). In addition, researches in the University of Hong Kong used Coronaviruses to hypothetically identify immunogenic parts of S protein [129].
4. Discussion
Rampant increase in the number of newly-identified patients with SARS-CoV-2 infection makes problematic challenges for occupational health managers, medical staff, economists, socialist, and governors [130, 131]. This highly-rapid spreading pandemic is deeply rooted in an improper recovery due to a poor/false diagnosis, and asymptomatic transmission. Totally, to globally curb long-term lung complications, diagnostic tests should be capable of rapid identification of SARS-CoV-2 infection [130, 132], and provide effective patient care among hospitalized patients. From immunopathological aspects, host immune responses of those critically-ill patients with COVID-19 (with pneumonia and severe lymphopenia), will proceed to Acute Respiratory Distress Syndrome (ARDS), shock, and death, necessitating a right frontline diagnostics in the shortest TAT, including laboratory-based diagnosis aimed at reducing falsifying results [130, 132, 133].
Accumulative studies declared that NAATs make a highly-sensitized viral detection by target genes, being majorly prioritized for clinical management of cases suspected with SARS-CoV-2 infection, directly measuring SARS-COV-2 nucleic acid by PCR-based techniques [134, 135]. Despite those mentioned advantages, as shortcomings, molecular diagnosis of SARS-CoV-2 infection cannot detect previous viral exposure, arising the interest of researchers to design clinical studies purposed to increase the sensitivity and specificity of those molecular tests [135-137]. Structurally, the key function of genetic material in Coronaviruses (a single-stranded, positive sense RNA) is encoded by the replicas gene, encoding two major polyproteins named pp1a and pp1ab. Hence, we can improve efficacy and sensitivity of the diagnostic procedures by simultaneous detection of two or more specific sequences through duplex or multiplex real-time RT-PCR test. Although, in case of accuracy, CRISPR and other lateral flow-based diagnostics are competing in a parallel manner with RT-PCR-based diagnosis [130, 133, 138, 139].
Using appropriately-stored samples from proper and various anatomic sites of cases suspected with SARS-CoV-2 infection accompanied with investigation of clinical features, can be effective in controlling viral spreading or disease aggravation, monitoring response to treatment, and assessment of viral infectivity [133, 137, 140]. Although existing viral RNA-based diagnostic tests are primarily qualitative, reliable results were reported from optimization of several synthetic RNAs in the process of UPE and ORF1b-based single-stage RT-PCR diagnostic approaches. Visiting the NCBI site with the code hCov-EmcJX869059 DNA UPE and ORF1b DNA Coronavirus patterns can lead to designation of specific primers and probes for the viral genome in PCR-based molecular testing [141, 142].
Although usage of appropriate samples in RT-PCR tests for cases suspected with SARS-CoV-2 infection have some limitation [143], repetitive (four times of) NP and OP sampling can definitely increase diagnostic sensitivity in mentioned cases upon admission to the hospitals [143].
Despite acceptable specificity and sensitivity of serologic-based tests for COVID-19 detection, false-negative results from reduced viral load after acute phase of disease, false-positive results due to the cross-reactivity of antibodies, and predisposition (autoimmune) diseases, lead us toward a combination of serological and molecular-based diagnosis to increase diagnostic sensitivity and specificity for cases suspected with SARS-CoV-2 infection [144-146].
There are validated diagnostic RT-PCR kits to detect viral presence in the biological samples [147]. As noted earlier, CT-based approaches can be practically recruited for reducing ambiguities in early clinical symptoms of COVID-19 patients, or asymptomatic carriers being exposured with clinically-diagnosed patients with COVID-19, or clinically-confirmed patients, or clinically-suspected cases with SARS-CoV-2 infection in the epidemic areas [94, 148-151]. As negative result of chest CT cannot exclude SARS-CoV-2 infection, clinicians and laboratory specialists should pay attention to the alternative/complementary strategies (repeatedly-over time test based on the assessment of each patient's clinical status) [152-154]. For instance, chest ultrasonography was reported for confirmation of positive results acquired from RT-PCR (being associated with Lung Ultrasound Score (LUSS), abnormal AST findings, and fever with an overall accuracy of 91%) in cases suspected with SARS-CoV-2 infection, aimed at reducing irradiation risks in CT-based approaches, and false-negativities in RT-PCR tests [155]. Moreover, triage-based strategies like fever in early screening of cases suspected with SARS-CoV-2 infection and negative results in their second nucleic acid test point out the establishment of fever clinics for reduction of misdiagnosis [156]. Moreover, Liver function damages, crazy-paving pattern, leukopenia, lymphopenia, elevated inflammatory factors can make indisputable roles for precise and rapid diagnosis aimed at early screening of cases suspected with SARS-CoV-2 infection [157-161].
Although, the role of CT for evaluation of cases suspected with COVID-19 infection is still yet to be specified [162], CT images can alternatively help differential diagnosis, early screening of COVID-19 or other pulmonary disorders, and initial evaluation of cases with non-specific clinical symptoms [162-165]. Due to lowered diagnostic specificity, it is worthy to mention that CT should not be used as a first-line screening approach for patients with COVID-19 and left-sided pneumonia [166], persistent indications of inflammation (despite receiving antibiotic regimens), repetitive negative results from NP swabs, and positive results from BAL samples in RT-PCR tests [166].
Accordingly, CT-based approaches can be initially of diagnostic specificity and quarantine necessity for cases suspected with SARS-CoV-2 infection and repetitive negativity in their RT-PCR tests on NP, OP, or BAL samples [155, 167]. Thereafter, combinative usage of chest CT and RT-PCR testing for febrile cases suspected with SARS-CoV-2 infection in EDs is recommended to reduce the possible false-negativities [168-172]. Additionally, COVID-19 Reporting and Data System (CO-RADS), as a categorical assessment scheme for prediction of infection rate and pulmonary involvement, increases specificity and diagnostic accuracy for cases suspected with SARS-CoV-2 infection [173-175].
5. Conclusion
According to the high risk of viral transmission from cases suspected with COVID-19, an accurate POC diagnostics can validate monitoring therapy of symptomatic patients, and prevent from other loaded nosocomial infections, as well, Correspondingly, “Precision/Personalized/Individualized Medicine”, as a missing piece in the puzzle of targeted diagnosis, can provide predictable outcomes for cases suspected with SARS-CoV-2 infection, and the most efficient vaccine with maximum immunogenicity with consideration of predisposition factors and immune-genetics of appropriate patients. Additionally, it remarkably minimizes negative predictive value of diagnostic tests, psychological burdens on health-care professionals, and exorbitant expenses of hospitalization imposed on patients. So, further pre-clinical investigations will be unquestionably needed to clarify the most efficient diagnostic protocol for SARS-CoV-2 infection. Moreover, scientific collaboration among specialists in the fields of internal/infectious diseases, pulmonary disorders, personalized medicine, immunology, medical microbiology/virology, medical biotechnology, medical genetics, medical laboratory, sciences, basic medical sciences, nursing, epidemiology, diseases-specific biomarkers, and health system coordinators is highly recommended.
Declaration Statement
Ethical Approval and Consent to Participate/for Publication
Not Applicable.
Author Contribution
Supervision and verification of the last version before submission, N.R.;
Conceptualization, Validation, and Formal analysis, N.R., and N.B.;
Main methodology, Search Strategy, and Data Curation, Academic/Scientific/Grammatical revision for important intellectual content, Preparation and main designation of the final draft of the manuscript, N.R., and N.B.;
Major Conception, Data extraction, Interpretation, and preparation of final sections related to clinical diagnosis of the study, M.A., and N.B.;
Preparation of first draft of manuscript related to clinical diagnosis of the study, M.A., S.S., N.M., and MJ.H.;
Major Conception, data extraction, Interpretation, and preparation of final sections related to laboratory diagnosis of the study, M.M., P.AE. SH.A. and N.B.;
Preparation of first draft of manuscript related to laboratory diagnosis of the study, M.M., P.AE., SH.A., F.R., E.N., P.F., M.M., A.M., H.K., M.SH., M.B., S.R., M.J., S.S., Z.D., S.L., S.M.M.K., SH.J., and S.E.;
Visualization, Image/Table Designation, N.B. and E.N.;
All of the authors attest to the validity and legitimacy of data, receiving an electronic copy of the final version, and published version of the manuscript.
Acknowledgements
No grant is received from public, commercial, or not-for-profit sectors funding agencies. All of the authors would like to sincerely dedicate this article to clinical specialists basic medical scientists, and health-care personnel all around the world, who are fighting with this pandemic situation as frontline agents.
Conflict of Interest
None
References
- Zhu N, et al., A novel coronavirus from patients with pneumonia in China, 2019. New England Journal of Medicine (2020).
- Fakheran O, Dehghannejad M, and Khademi A. Saliva as a diagnostic specimen for detection of SARS-CoV-2 in suspected patients: a scoping review. Infectious diseases of poverty 9 (2020): 1-7.
- Gorbalenya AE, et al., Severe acute respiratory syndrome-related coron-avirus: The species and its viruses–a statement of the Coronavirus Study Group (2020).
- Chi Q, et al., Differential diagnosis for suspected cases of coronavirus disease 2019: a retrospective study. BMC Infectious Diseases (2020).
- Xie S, et al., The Epidemiologic and Clinical Features of Suspected and Confirmed Cases of Imported 2019 Novel Coronavirus Pneumonia in North Shanghai, China. China (2020).
- Jahromi AH and Mahmoudi H, Estimates of mortality following COVID-19 Infection; comparison between Europe and the United States. Immunopathologia Persa 7 (2020): e05-e05.
- Anka AU, et al, Coronavirus Disease 2019 (COVID-19): An Overview of the Immunopathology, Serological Diagnosis and Management. Scandinavian Journal of Immunology (2021): e12998.
- Araujo MB, and Naimi B, Spread of SARS-CoV-2 Coronavirus likely to be constrained by climate. medRxiv (2020).
- Lippi G. Brief update on coronavirus disease 2019 (COVID-19) diagnostics. Advances in Laboratory Medicine-/Avances en Medicina de Laboratorio (2020).
- Wang Y-Y, et al., Updating the diagnostic criteria of COVID-19 “suspected case” and “confirmed case” is necessary. Military Medical Research 7 (2020): 1-3.
- Bennett CE, et al. ST-segment elevation, myocardial injury, and suspected or confirmed COVID-19 patients: diagnostic and treatment uncertainties. in Mayo Clinic Proceedings (2020).
- Eljilany I, and A-N. Elzouki, D-Dimer, Fibrinogen, and IL-6 in COVID-19 Patients with Suspected Venous Thromboembolism: A Narrative Review. Vascular health and risk management 16 (2020): 455.
- Hanley B, et al., Autopsy in suspected COVID-19 cases. Journal of clinical pathology 73 (2020): 239-242.
- Daneshfar M, et al., Lessons of mortality following COVID-19 epidemic in the United States especially in the geriatrics. Journal of Nephropharmacology 10 (2021): 1-4.
- Kampf G, et al., Persistence of coronaviruses on inanimate surfaces and their inactivation with biocidal agents. Journal of Hospital Infection 104 (2020): 246-251.
- Sun G, et al., Clinical features and outcomes of pregnant women suspected of coronavirus disease 2019. Journal of Infection (2020).
- Moradian N, et al., The urgent need for integrated science to fight COVID-19 pandemic and beyond. Journal of Translational Medicine 18 (2020): 1-7.
- Moazzami B, et al., Is computed tomography necessary for the diagnosis of coronavirus disease (Covid–19) in all suspected patients? a case series. Journal of Kerman University of Medical Sciences 28 (2021): 187-193.
- Dana Ghotbi, Elham Nouri, Nazila Bahmaie, Abdolreza Esmaeilzadeh. Coronavirus Diagnosis: Time for Unravelling the Secrets and Science Without Boundaries. Journal of Emerging Diseases and Preventive Medicine 3 (2020).
- Cao G, et al., The potential transmission of SARS-CoV-2 from patients with negative RT-PCR swab tests to others: two related clusters of COVID-19 outbreak. Japanese Journal of Infectious Diseases, 165 (2020): JJID.
- Chua F, et al., The role of CT in case ascertainment and management of COVID-19 pneumonia in the UK: insights from high-incidence regions. The Lancet Respiratory Medicine 8 (2020): 438-440.
- Makurumidze R. Coronavirus-19 Disease (COVID-19): a case series of early suspects reported and the implications towards the response to the pandemic in Zimbabwe. Journal of Microbiology, Immunology and Infection (2020).
- da Silva SJR, et al, Clinical and laboratory diagnosis of SARS-CoV-2, the virus causing COVID-19. ACS infectious diseases 6 (2020): 2319-2336.
- Zheng H, et al., Perioperative management of patients with suspected or confirmed COVID-19: review and recommendations for perioperative management from a retrospective cohort study. British journal of anaesthesia (2020).
- Lubega M and Ekol JE. Preparing communities to receive persons recently suspected or diagnosed with COVID-19. The Pan African Medical Journal 35 (2020).
- Banerjee A, et al., Excess deaths in people with cardiovascular diseases during the COVID-19 pandemic. European journal of preventive cardiology (2020).
- Saghafi AM, Aghaali, and Saghafi H. Acute kidney injury in hospitalized COVID-19 patients in Iran; a systematic review and meta-analysis. Journal of Renal Injury Prevention 10 (2021): 09-09.
- Onuigbo MAC. Persistence of coronavirus disease 2019 (COVID-19) in patients with end-stage renal disease; an unrecognized phenomenon? Journal of Renal Injury Prevention 10 (2020): e07-e07.
- Sadighpour TMM, Sabaeifard P, Saeifar S, Kenari F. COVID-19 and renal involvement; evolving role of thromboinflammation, vascular and glomerular disease in the pathogenesis. Nephropathology 10 (2021).
- Gharehbaghi G, et al., COVID-19 in Children and Neonates: A Comprehensive Review Article. Iranian Journal of Pediatrics 31 (2021).
- Aleebrahim-Dehkordi E, et al., Human coronaviruses SARS-CoV, MERS-CoV, and SARS-CoV-2 in children. Journal of Pediatric Nursing (2020).
- Sarzaeim M and Rezaei N. Kawasaki Disease and Multisystem Inflammatory Syndrome in Children with COVID-19. SN comprehensive clinical medicine (2020): 1-6.
- Hessami A, et al., Cardiovascular diseases burden in COVID-19: Systematic review and meta-analysis. The American journal of emergency medicine (2020).
- Mizumoto K, et al., Estimating the asymptomatic proportion of coronavirus disease 2019 (COVID-19) cases on board the Diamond Princess cruise ship, Yokohama, Japan, 2020. Eurosurveillance 25 (2020): 2000180.
- Chen X, et al., Differences between COVID-19 and suspected then confirmed SARS-CoV-2-negative pneumonia: a retrospective study from a single center. Journal of Medical Virology (2020).
- Besutti G, et al., Accuracy of CT in a cohort of symptomatic patients with suspected COVID-19 pneumonia during the outbreak peak in Italy. European radiology (2020): 1-10.
- Santurro A, et al, A technical report from the Italian SARS-CoV-2 outbreak. Postmortem sampling and autopsy investigation in cases of suspected or probable COVID-19. Forensic Science, Medicine, and Pathology (2020): 1.
- Shahzad N, et al., Rapid assessment of COVID-19 suspected cases: A community based approach for developing countries like Pakistan. Journal of Global Health 10 (2020).
- Lumba R, et al., Neonate Born to a Mother with a Diagnosis of Suspected Intra-Amniotic Infection versus COVID-19 or Both. Case Reports in Pediatrics 2020 (2020).
- Peng D, et al., Clinical features of suspected pediatric patients with 2019 novel coronavirus infection and the role of procalcitonin in early differential diagnosis. medRxiv (2020).
- Leal FE, et al., A primary care approach to the COVID-19 pandemic: clinical features and natural history of 2,073 suspected cases in the Corona Sao Caetano programme, Sao Paulo, Brazil. medRxiv (2020).
- Prabhakaran D et al., Management of cardiovascular disease patients with confirmed or suspected COVID-19 in limited resource settings. Global heart 15 (2020).
- Amrane S, et al., Rapid viral diagnosis and ambulatory management of suspected COVID-19 cases presenting at the infectious diseases referral hospital in Marseille, France,-January 31st to March 1st, 2020: A respiratory virus snapshot. Travel medicine and infectious disease (2020): 101632.
- Silva EH and Lekamwasam S, Confirmatory and supportive laboratory investigations in SARS-CoV-2 infection; a systematic review. Journal of Preventive Epidemiology 5 (2020): e03-e03.
- Alteri C, et al., Detection and quantification of SARS-CoV-2 by droplet digital PCR in real-time PCR negative nasopharyngeal swabs from suspected COVID-19 patients. PloS one 15 (2020): e0236311.
- Di Paolo M, et al, False-negative RT-PCR in SARS-CoV-2 disease: experience from an Italian COVID-19 unit. ERJ open research 6 (2020).
- Gao X, et al, Improving the early diagnosis of suspected patients with COVID-19: a retrospective study of 106 patients. The Journal of Infection in Developing Countries 14 (2020): 547-553.
- Brendish NJ, et al, Clinical impact of molecular point-of-care testing for suspected COVID-19 in hospital (COV-19POC): a prospective, interventional, non-randomised, controlled study. The Lancet Respiratory Medicine (2020).
- Fang FC, Naccache SN, and Greninger AL, The Laboratory Diagnosis of COVID-19--Frequently-Asked Questions. Clinical Infectious Diseases (2020).
- Luo L, et al, CT differential diagnosis of COVID-19 and non-COVID-19 in symptomatic suspects: a practical scoring method. BMC Pulmonary Medicine 20 (2020): 1-9.
- Nakakubo S, et al., Proposal of COVID-19 Clinical Risk Score for the management of suspected COVID-19 cases: a case control study. BMC Infectious Diseases 20 (2020): 1-11.
- Sampaio SGdSM, et al., Evaluation of the Criteria Adopted to Identify Suspected Cases of COVID-19 in the Emergency Department Service of a Referral Palliative Oncology Care Unit. American Journal of Hospice and Palliative Medicine® (2020): 1049909120957310.
- Carter LJ, et al., Assay Techniques and Test Development for COVID-19 Diagnosis. ACS Publications (2020).
- Santantonio TA. and Messina G, Update on Coronavirus Disease 2019 (COVID-19). The Open Neurology Journal 14 (2020).
- Martin-Sanchez FJ, et al., Diagnostic groups and short-term outcomes in suspected COVID-19 cases treated in an emergency department. Emergencias: revista de la Sociedad Espanola de Medicina de Emergencias (2020).
- Islam N, et al., Thoracic imaging tests for the diagnosis of COVID-19. Cochrane Database of Systematic Reviews (2021).
- Mironas A et al., How to best test suspected cases of COVID-19: an analysis of the diagnostic performance of RT-PCR and alternative molecular methods for the detection of SARS-CoV-2. medRxiv (2021).
- Abasiyanik MF, et al., Sensitive detection and quantification of SARS-CoV-2 in saliva. Scientific reports 11 (2021): 1-12.
- Deeks JJ et al., Diagnosis of SARS-CoV-2 infection and COVID-19: accuracy of signs and symptoms; molecular, antigen, and antibody tests; and routine laboratory markers. Cochrane Database of Systematic Reviews (2020).
- Iser BPM, et al., Suspected COVID-19 case definition: a narrative review of the most frequent signs and symptoms among confirmed cases. Epidemiologia e Serviços de Saúde 29 (2020): e2020233.
- Xu K et al, Management of corona virus disease-19 (COVID-19): the Zhejiang experience. Journal of Zhejiang University (medical science) 49 (2020): 0-0.
- Zhang J, Wang S, and Xue Y, Fecal specimen diagnosis 2019 novel coronavirus–infected pneumonia. Journal of medical virology 92 (2020): 680-682.
- Zhang Y, et al, Isolation of 2019-nCoV from a stool specimen of a laboratory-confirmed case of the coronavirus disease 2019 (COVID-19). China CDC Weekly 2 (2020): 123-124.
- Xie C, et al., Comparison of different samples for 2019 novel coronavirus detection by nucleic acid amplification tests. International Journal of Infectious Diseases (2020).
- Kim D, et al., Rates of co-infection between SARS-CoV-2 and other respiratory pathogens. Jama (2020).
- Organization WH. Infection prevention and control during health care when novel coronavirus ( nCoV) infection is suspected: interim guidance, 25 January 2020 (2020).
- Shi X, et al. Severe acute respiratory syndrome associated coronavirus is detected in intestinal tissues of fatal cases. American Journal of Gastroenterology 100 (2005): 169-176.
- Rodriguez-Morales AJ, et al., Clinical, laboratory and imaging features of COVID-19: A systematic review and meta-analysis. Travel medicine and infectious disease (2020): 101623.
- Ishige T, et al. Highly sensitive detection of SARS-CoV-2 RNA by multiplex rRT-PCR for molecular diagnosis of COVID-19 by clinical laboratories. Clinica Chimica Acta (2020).
- Soraya GV. and Ulhaq ZS. Crucial Laboratory Parameters in COVID-19 Diagnosis and Prognosis: An Updated Meta-Analysis. Available at SSRN 3576912 (2020).
- Yip CC-Y, et al. Development of a Novel, Genome Subtraction-Derived, SARS-CoV-2-Specific COVID-19-nsp2 Real-Time RT-PCR Assay and Its Evaluation Using Clinical Specimens. International Journal of Molecular Sciences 21 (2020): 2574.
- Sharma R, et al., Clinical Characteristics and Differential Clinical Diagnosis of Novel Coronavirus Disease 2019 (COVID-19), in Coronavirus Disease 2019 (COVID-19) (2020): 55-70.
- Sheridan C, Fast, portable tests come online to curb coronavirus pandemic. Nat Biotechnol 10 (2020).
- Tahamtan, A. and A. Ardebili, Real-time RT-PCR in COVID-19 detection: issues affecting the results (2020).
- Yelin I, et al. Evaluation of COVID-19 RT-qPCR test in multi-sample pools. medRxiv (2020).
- Zhong L, et al. Detection of serum IgM and IgG for COVID-19 diagnosis. Science China Life sciences 63 (2020): 777-780.
- Broughton, J.P., et al., CRISPR–Cas12-based detection of SARS-CoV-2. Nature Biotechnology (2020): 1-5.
- Broughton JP, et al. Rapid detection of 2019 novel coronavirus SARS-CoV-2 using a CRISPR-based DETECTR lateral flow assay. MedRxiv (2020).
- Joynt GM and Wu WK, Understanding COVID-19: what does viral RNA load really mean? The Lancet Infectious Diseases (2020).
- Lan L et al. Positive RT-PCR test results in patients recovered from COVID-19. Jama 323 (2020): 1502-1503.
- Sethuraman N, Jeremiah SS and Ryo A, Interpreting Diagnostic Tests for SARS-CoV-2. JAMA (2020).
- Long C, et al. Diagnosis of the Coronavirus disease (COVID-19): rRT-PCR or CT? European journal of radiology (2020): 108961.
- Zhou J, et al. Human intestinal tract serves as an alternative infection route for Middle East respiratory syndrome coronavirus. Science advances 3 (2017): eaao4966.
- Ding Y, et al. Organ distribution of severe acute respiratory syndrome (SARS) associated coronavirus (SARS-CoV) in SARS patients: implications for pathogenesis and virus transmission pathways. The Journal of Pathology: A Journal of the Pathological Society of Great Britain and Ireland 203 (2004): 622-630.
- Organization WH. Guidance on regulations for the transport of infectious substances: applicable from 1 January 2019. 2019, World Health Organization (2019–2020).
- Lippi GHB, Sanchis-Gomar F, Mattiuzzi C, Updates on laboratory investigations in coronavirus disease 2019 (COVID-19). Acta Bio Med 91 (2020).
- Zhang Jj, et al, Distinct characteristics of COVID-19 patients with initial rRT-PCR-positive and rRT-PCR-negative results for SARS-CoV-2. Allergy (2020).
- Bordi L, et al, Differential diagnosis of illness in patients under investigation for the novel coronavirus (SARS-CoV-2), Italy, February 2020. Eurosurveillance 25 (2020): 2000170.
- Iyer M et al. COVID-19: an update on diagnostic and therapeutic approaches. BMB reports 53 (2020): 191-205.
- Organization WH. Laboratory testing for coronavirus disease ( COVID-19) in suspected human cases: interim guidance, 19 March 2020. World Health Organization (2020).
- Organization WH. Laboratory testing for coronavirus disease 2019 (COVID-19) in suspected human cases: interim guidance, 2 March 2020.World Health Organization (2020).
- Patel R, et al. Report from the american society for microbiology covid-19 international summit, 23 march 2020: Value of diagnostic testing for sars–cov-2/covid-19. Am Soc Microbiol (2020).
- He J-L, et al. Diagnostic performance between CT and initial real-time RT-PCR for clinically suspected 2019 coronavirus disease (COVID-19) patients outside Wuhan, China. Respiratory Medicine (2020): 105980.
- Zhu W, et al. Initial clinical features of suspected coronavirus disease 2019 in two emergency departments outside of Hubei, China. Journal of Medical Virology (2020).
- Datta A, et al, Evaluation of Diagnostic Performance of Chest CT with RT-PCR in Suspected Coronavirus Patients in Bangladesh. Mymensingh Medical Journal: MMJ 29 (2020): 964-968.
- Kuzan TY, et al. A comparison of clinical, laboratory and chest CT findings of laboratory-confirmed and clinically diagnosed COVID-19 patients at first admission. Diagnostic and Interventional Radiology (Ankara, Turkey) (2020).
- Xie S, et al. Chest CT-based differential diagnosis of 28 patients with suspected corona virus disease 2019 (COVID-19). The British Journal of Radiology 93 (2020): 20200243.
- Miao C, et al. Early chest computed tomography to diagnose COVID-19 from suspected patients: A multicenter retrospective study. The American Journal of Emergency Medicine (2020).
- Harkin, T.J., et al., Delayed diagnosis of COVID-19 in a 34-year-old man with atypical presentation. The Lancet. Respiratory Medicine (2020).
- Hao Q, Wu H and Wang Q, Difficulties in False Negative Diagnosis of Coronavirus Disease 2019: A Case Report. (2020).
- Khodamoradi Z, Moghadami M, and Lotfi M, Co-infection of coronavirus disease 2019 and influenza: a report from Iran. Archives of Iranian Medicine 23 (2020): 239-243.
- Hornuss D, et al. COVID-19 associated pneumonia despite repeatedly negative PCR-analysis from oropharyngeal swabs. Deutsche medizinische Wochenschrift (1946) 145 (2020): 844-849.
- Chen L-D, et al. A COVID-19 patient with multiple negative results for PCR assays outside Wuhan, China: a case repor. (2020).
- Hase R, et al. A case of imported COVID-19 diagnosed by PCR-positive lower respiratory specimen but with PCR-negative throat swabs. Infectious Diseases 52 (2020): 423-426.
- Giannitto C, et al. Chest CT in patients with a moderate or high pretest probability of COVID-19 and negative swab. La radiologia medica 125 (2020): 1260-1270.
- Lin C, et al. Comparison of throat swabs and sputum specimens for viral nucleic acid detection in 52 cases of novel coronavirus (SARS-Cov-2)-infected pneumonia (COVID-19). Clinical Chemistry and Laboratory Medicine (CCLM) 58 (2020): 1089-1094.
- Poortahmasebi V, et al. Clinical performance of RT-PCR and chest CT scan for COVID-19 diagnosis; a systematic review. Advanced Journal of Emergency Medicine 4 (2020): e57-e57.
- Yang D, et al. A suspected case of COVID-19 turned into a confirmed case: a case report from Iran. Future Virology (2020).
- Yang D, et al. A suspected case of COVID-19 turned into a confirmed case: a case report. Future Virology 15 (2020): 335-339.
- Habli Z, et al. COVID-19 in-vitro diagnostics: state-of-the-art and challenges for rapid, scalable, and high-accuracy screening. Frontiers in bioengineering and biotechnology 8 (2021): 1562.
- Meyer B, Drosten C, and Müller MA, Serological assays for emerging coronaviruses: challenges and pitfalls. Virus research 194 (2014): 175-183.
- Zeng H, et al. Diagnostic value of combined nucleic acid and antibody detection in suspected COVID-19 cases. Public health 186 (2020): 1-5.
- Long Q-X. et al. Antibody responses to SARS-CoV-2 in patients with COVID-19. Nature medicine (2020): 1-4.
- Li Z, et al. Development and clinical application of a rapid IgM-IgG combined antibody test for SARS-CoV-2 infection diagnosis. Journal of medical virology (2020).
- Lin D, et al. Evaluations of serological test in the diagnosis of 2019 novel coronavirus (SARS-CoV-2) infections during the COVID-19 outbreak. medRxiv (2020).
- Shen B, et al. Clinical evaluation of a rapid colloidal gold immunochromatography assay for SARS-Cov-2 IgM/IgG. American journal of translational research 12 (2020): 1348.
- Infantino M, et al. Diagnostic accuracy of an automated chemiluminescent immunoassay for anti-SARS-CoV-2 IgM and IgG antibodies: an Italian experience. Journal of medical virology (2020).
- Ksiazek TG, et al. A novel coronavirus associated with severe acute respiratory syndrome. New England journal of medicine 348 (2003): 1953-1966.
- Lippi G and Plebani M, Laboratory medicine resilience during coronavirus disease 2019 (COVID-19) pandemic. Advances in Laboratory Medicine/Avances en Medicina de Laboratorio 1 (2020).
- To KK-W, et al. Temporal profiles of viral load in posterior oropharyngeal saliva samples and serum antibody responses during infection by SARS-CoV-2: an observational cohort study. The Lancet Infectious Diseases (2020).
- Ings DP, et al. Few SARS-CoV-2 infections detected in Newfoundland and Labrador in the absence of Public Health Laboratory-based confirmation. Plos one 17 (2022): e0262957.
- Feng Y. Clinical Value of SARS-CoV2 IgM and IgG Antibodies in Diagnosis of COVID-19 in Suspected Cases. Journal of Inflammation Research 13 (2020): 1089.
- Jia X, et al. Clinical significance of IgM and IgG test for diagnosis of highly suspected COVID-19 infection. medRxiv (2020).
- Xiang F, et al. Antibody detection and dynamic characteristics in patients with COVID-19. Clinical Infectious Diseases (2020).
- Pizzol JLD, et al. Laboratory diagnosis for Covid-19: A mini-review. Revista da Sociedade Brasileira de Medicina Tropical 53 (2020).
- Shaoli B and Jianyun W, Zhou (Yingquan, Yu Desheng, Gao Xiaomin, Li Lingling, Yang Fan. Analysis of the first family epidemic situation of new coronavirus pneumonia in Gansu Province. Chinese Journal of Preventive medicine 54 (2020).
- Iwasaki A. and Yang Y, The potential danger of suboptimal antibody responses in COVID-19. Nature Reviews Immunology (2020): 1-3.
- Stefanelli P, et al. Whole genome and phylogenetic analysis of two SARS-CoV-2 strains isolated in Italy in January and February 2020: additional clues on multiple introductions and further circulation in Europe. Eurosurveillance 25 (2020): 2000305.
- Organization WH. Joint external evaluation tool: International Health Regulations ( 2005). (2018).
- Afzal A. Molecular diagnostic technologies for COVID-19: Limitations and challenges. Journal of advanced research (2020).
- Hanson KE, et al, The Infectious Diseases Society of America Guidelines on the Diagnosis of COVID-19: Molecular Diagnostic Testing. infect. dis (2021).
- Syal K. Guidelines on newly identified limitations of diagnostic tools for COVID-19 and consequences. Journal of medical virology 93 (2021): 1837-1842.
- Islam KU and Iqbal J. An Update on Molecular Diagnostics for COVID-19. Frontiers in cellular and infection microbiology (2020).
- Falzone L, et al. Current and innovative methods for the diagnosis of COVID-19 infection. International journal of molecular medicine 47 (2021): 1-23.
- Zavvar M, et al. SARS-Cov-2 and COVID-19, Basic and Clinical Aspects of the Human Pandemic: A Review. Iranian Journal of Public Health 50 (2021): 665-675.
- Organization WH. Protocol to investigate non-seasonal influenza and other emerging acute respiratory diseases. World Health Organization (2018).
- Feng W, et al. Molecular diagnosis of COVID-19: challenges and research needs. Analytical chemistry 92 (2020): 10196-10209.
- Zhou P, et al. Discovery of a novel coronavirus associated with the recent pneumonia outbreak in humans and its potential bat origin. BioRxiv (2020).
- Rahimi H, et al. CRISPR systems for COVID-19 diagnosis. ACS sensors 6 (2021): 1430-1445.
- Oishee MJ, et al., COVID-19 pandemic: review of contemporary and forthcoming detection tools. Infection and drug resistance 14 (2021): 1049.
- Gomes JC, et al. Optimizing the molecular diagnosis of Covid-19 by combining RT-PCR and a pseudo-convolutional machine learning approach to characterize virus DNA sequences. bioRxiv (2020).
- Wang Y, et al, Combination of RT-qPCR testing and clinical features for diagnosis of COVID-19 facilitates management of SARS-CoV-2 outbreak. Journal of medical virology (2020).
- Lee KK, et al., Diagnostic performance of the combined nasal and throat swab in patients admitted to hospital with suspected COVID-19. BMC infectious diseases 21 (2021): p. 1-11.
- Liu G and Rusling JF, COVID-19 antibody tests and their limitations. ACS sensors 6 (2021): 593-612.
- Peeling RW, et al. Scaling up COVID-19 rapid antigen tests: promises and challenges. The Lancet infectious diseases, 2021.
- Dinnes J, et al, Rapid, point-of-care antigen and molecular-based tests for diagnosis of SARS-CoV-2 infection. Cochrane Database of Systematic Reviews 3 (2021).
- Nelson AC, et al, Analytical Validation of a COVID-19 qRT-PCR Detection Assay Using a 384-well Format and Three Extraction Methods. BioRxiv (2020).
- Pakdemirli E, Mandalia U, and Monib S, Positive chest CT features in patients with COVID-19 pneumonia and negative real-time polymerase chain reaction test. Cureus 12 (2020).
- De Smet K, et al. Diagnostic performance of chest CT for SARS-CoV-2 infection in individuals with or without COVID-19 symptoms. Radiology (2020): 202708.
- Majidi H and Niksolat F, Chest CT in patients suspected of COVID-19 infection: A reliable alternative for RT-PCR. The American Journal of Emergency Medicine (2020).
- Chen D, et al. Can Chest CT Features Distinguish Patients With Negative From Those With Positive Initial RT-PCR Results for Coronavirus Disease (COVID-19)? American Journal of Roentgenology (2020): 1-5.
- Dramé M, et al. Should RT-PCR be considered a gold standard in the diagnosis of Covid-19? Journal of Medical Virology (2020).
- Krdzalic JTM. de Jaegere and Kwee RM, Diagnostic performance of chest CT in screening patients with suspected COVID-19 infection in a Western population. The British journal of radiology 93 (2020): p. 20200643.
- Wang S, et al. A deep learning algorithm using CT images to screen for Corona Virus Disease (COVID-19). European radiology (2021): p. 1-9.
- Allegorico E, et al, The use of chest ultrasonography in suspected cases of COVID-19 in the emergency department. Future Science OA 7 (2020): p. FSO635.
- Jia H, et al, The Role of Fever Clinics in the Strategic Triage of Suspected Cases of Imported COVID-19. International Journal of General Medicine 14 (2021): p. 2047.
- Stegeman I, et al, Routine laboratory testing to determine if a patient has COVID-19. Cochrane Database of Systematic Reviews 2020(11).
- Mardani R, et al. Laboratory parameters in detection of COVID-19 patients with positive RT-PCR; a diagnostic accuracy study. Archives of academic emergency medicine 8 (2020).
- Miao C, et al. A comparative multi-centre study on the clinical and imaging features of comfirmed and uncomfirmed patients with COVID-19. MedRxiv (2020).
- Zhao D, et al, comparative study on the clinical features of COVID-19 pneumonia to other pneumonias; Clinical Infectious Diseases; Oxford Academic. Clinical Infectious Diseases.
- Yang Z, et al, Distinguishing COVID-19 from influenza pneumonia in the early stage through CT imaging and clinical features. medRxiv (2020).
- Elicker BM, What is the Performance and Role of CT in Suspected COVID-19 Infection? 2021, Radiological Society of North America.
- Nezhad MS, et al. An overview of the prominence of current diagnostic methods for diagnosis of COVID-19. AIMS Allergy and Immunology 4 (2020): p. 60-74.
- Abosamak MF, et al. Is There a Role for Pre-Operative CT Chest Scans in Suspected COVID-19 Patients Requiring Emergent Surgery?: Insights from a Cross-Sectional Study. Insights from a Cross-Sectional Study (March 11, 2021) (2021).
- Wu L, et al. Diagnostic accuracy of chest computed tomography scans for suspected patients with COVID-19: Receiver operating characteristic curve analysis. JMIR public health and surveillance 6 (2020): e19424.
- Chrzan R, et al. COVID-19 Infection Negative in Nasopharyngeal Swabs but Suspected in Computed Tomography and Confirmed in Bronchoalveolar Lavage Material. Case Reports in Infectious Diseases (2021).
- Salehi-Pourmehr H, et al. Diagnostic value of chest CT in Iranian patients with suspected COVID-19. Caspian Journal of Internal Medicine 11 (2020): 527.
- Korkmaz I, et al. Chest CT in COVID-19 pneumonia: correlations of imaging findings in clinically suspected but repeatedly RT-PCR test-negative patients. Egyptian Journal of Radiology and Nuclear Medicine 52 (2021): p. 1-9.
- Korevaar DA, et al. Added value of chest computed tomography in suspected COVID-19: an analysis of 239 patients. European Respiratory Journal 56 (2020).
- Cartocci G, et al. Correction to: Chest CT for early detection and management of coronavirus disease (COVID-19): a report of 314 patients admitted to Emergency Department with suspected pneumonia (La radiologia medica 125 (2020): 931-942.
- Song S, et al. Correlation between chest CT findings and clinical features of 211 COVID-19 suspected patients in Wuhan, China. in Open forum infectious diseases. (2020)
- De Smet K, et al, Diagnostic power of chest CT for COVID-19: to screen or not to screen. Available at SSRN 3605168 (2020).
- Fujioka T, et al. Evaluation of the usefulness of CO-RADS for chest CT in patients suspected of having COVID-19. Diagnostics 10 (2020): 608.
- Prokop M, et al. CO-RADS: a categorical CT assessment scheme for patients suspected of having COVID-19—definition and evaluation. Radiology 296 (2020): E97-E104.
- Bellini D, et al. Diagnostic accuracy and interobserver variability of CO-RADS in patients with suspected coronavirus disease-2019: a multireader validation study. European radiology 31 (2021): 1932-1940.