Characterization and use of Helicobacter pylori SecAN68 ATPase for probing SecA inhibitors as antibacterial agents
Article Information
Han, Qiong 1, Tiantian Jian1, Yan Liu1, Han Li1, Junda Zhou1, Hyuk-kyu Seoh2, Sen-Fang Sui3, Xu Jia4, Phang-Cheng Tai2, *, Xinhe Huang1, *
Affiliation:
1School of Life Sciences and Engineering, Southwest Jiaotong University, Chengdu 610031, China.
2Department of Biology, Georgia State University, Atlanta, GA 30303, USA
3State Key Laboratory of Membrane Biology, Center
for Structural Biology, School of Life Sciences, Tsinghua University, Beijing 100084 China.
4Noncoding RNA and Drugs Key Laboratory of Sichuan Province, Chengdu Medical College, Chengdu 610599, China.
*Corresponding author:
Xinhe Huang, School of Life Sciences and Engineering, Southwest Jiaotong University, Chengdu 610031, China.
Phang-Cheng Tai, Department of Biology, Georgia State University, Atlanta, GA 30303, USA.
Received: November 22, 2023; Accepted: November 29, 2023; Published: December 13, 2023
Citation: Han, Qiong, Tiantian Jian, Yan Liu, Han Li, Junda Zhou, Hyuk-kyu Seoh, Sen-Fang Sui, Xu Jia, Phang-Cheng Tai, Xinhe Huang. Characterization and use of Helicobacter pylori SecAN68 ATPase for probing SecA inhibitors as antibacterial agents. Archives of Microbiology and Immunology. 7 (2023): 421-427.
Share at FacebookAbstract
SecA, the only known energy conversion ATPase in the bacterial protein transport pathway, is a unique protein not found in mammalian cells. H. pylori SecA is closely related to the secretion of the virulence factor vacuolating toxin (VacA) in mediating H. pylori infection and diseases. We previously identified several potential compounds through virtual screening that may inhibit SecA function. In this study for testing inhibitory activity, the HpSecAN68 fragments were constructed and cloned into pET-28a (+) to obtain recombinant plasmid for transforming into E. coli BL21 (DE3). NI-Nitrilotriacetic acid resins were employed to purify the HpSecAN68 protein that has high intrinsic ATPase activity. The recombinant HpSecAN68 protein was successfully expressed, optimized and purified in an active soluble form. The ATPase activity assay system using NADH colorimetry for the purified HpSecAN68 protein was established. The determined Km and Vmax of HpSecAN68 are 2.997±0.697 mM and 0.231±0.038 μmol·mg-1·min-1 respectively in the reaction system. The purified HpSecAN68 ATPase system was used to probe the inhibitory effects of three small molecule inhibitors identified in our study, two of these have IC50 values about 10 μM. In addition, using the micro-broth method, we also observed the significant antibacterial growth effects of these molecules for three Gram-positive and Gram-negative bacteria. This study provides a solid foundation for further research on H. pylori SecA and the development of SecA inhibitors as antimicrobial targets.
Keywords
H. pylori; SecA; ATPase properties; kinetic parameter, small molecules SecA inhibitors, antibacterial agents
H. pylori articles; SecA articles; ATPase properties articles; kinetic parameter articles, small molecules SecA inhibitors articles, antibacterial agents articles
Article Details
Introduction
Helicobacter pylori (H. pylori) is a Gram-negative micro-anaerobic pathogenic bacterium. H. pylori infection can cause many diseases including peptic ulcers, chronic gastritis, and gastric cancer. According to the World Gastroenterology Organization (WGO) data, more than 50% of the world's population has been infected with H. pylori. In countries like China, the H. pylori-carrying population is as high as 65% along with a high incidence of pylori infection and gastric cancer because of unhealthy eating habits [1]. At present, "quadruple therapy" (including at least two antibiotics) is used to treat H. pylori infection, however, with the wide-spread emergence of drug resistance, which leads to at least 10 million people dying [2]. Long- term medical practice shows that the weaknesses and ineffectiveness of "quadruple therapy" are gradually exposed. Therefore, it is imminent to develop new anti-H. pylori inhibitors whose mechanism of action is different from that of existing antibiotics. SecA is a key member of the AAA family of ATPases in harnessing the energy of ATP hydrolysis for mechanical work and guiding protein transmembrane transport, degradation, and unfolding [3]. SecA moves precursor substrates with signal peptides across bacterial membranes through the conduction protein SecYEG-dependent channel, which consists of three main polypeptide chains (SecY, SecE, and SecG) [4] and SecDFC, or the less efficient SecYEG-independent SecA- alone channels [5]. SecA protein is a multi-domain protein with RecA-like domains (NBD1 and NBD2) binding to SecYEG, phospholipid bilayers, and ribosomes [6]. Bacterial precursors could be translocated co- or post-translationally across membranes [5, 7]. After the signal peptides interaction, the SecA protein switches from the ADP-bound state to the ATP-bound state, and the SecYEG channel opens, allowing the peptide residues to diffuse through the pore. After ATP is hydrolyzed, the channel closes, trapping a large number of amino acid residues on the other side of the membrane, thus completing the protein transport [8]. Recent studies have shown that, in addition to SecYEG translocation channels, SecA-only channels exist in lipid bilayers that are functionally independent of the SecYEG mechanism [9,10,11]. Compared to the high-affinity SecYEG channel, the SecA-only channel is the original low-affinity channel with lower specificity but can function normally in the absence of SecYEG [11]. Current research shows that SecA acts as an "energy pump" in the Sec protein transport pathway, driving peptides through the channel via the hydrolysis cycle of ATP [12]. Since SecA protein is only found in bacteria, but not in eukaryotic cells, the development of inhibitors and bactericidal agents based on SecA protein is becoming a hot topic in this field recently. It has been shown that SecA is involved in the translocation of H. pylori vacuolar toxin (VacA), and the inhibition of SecA synthesis significantly reduces VacA secretion, which is greatly involved in the pathogenesis of H. pylori [13-16]. So SecA can be used as an excellent target for anti-H. pylori action. H. pylori SecA monomer has a molecular weight of 99 kDa and contains three basic domains: the dead driver, the substrate-specific binding region, and the C-terminal domain (34 kDa), though the crystal structure of H. pylori SecA has not yet been resolved. It has been shown that the C-terminal domain of the SecA protein has an inhibitory regulatory effect on ATPase activity, and the E. coil SecAN68 fragment can act as a high intrinsic ATPase substrate for testing SecA inhibitors [17]. In this study, we constructed a pET-28a (+)-HpSecAN68 recombinant plasmid and obtained a highly expressed and soluble recombinant H. pylori SecAN68 (HpSecAN68) protein with good catalytic ATPase activity. We measured the inhibition of ATP activity of three different pylori SecA small molecule inhibitors identified by virtual screening in our previous study [18], and obtained IC50 at the enzyme level. These SecA inhibitors also showed significant bacterial growth inhibition. This study laid a solid foundation for further development for optimization of inhibitors against pylori and other bacteria.
Materials and Methods
Materials
Plasmids, Strains, and Reagents are shown in Supplementary Information.
Methods
Construction of recombinant strains, Expression of recombinant plasmids, Optimization of expression conditions, Protein purification and the BCA Protein Test Kit used to determine the content of purified HpSecAN68, the methods and experimental results details are shown in Supplementary Information.
Analysis of the ATPase enzymatic properties and determination of kinetic parameters of recombinant HpSecAN68
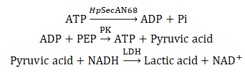
SecA ATPase catalyzes the production of ADP and phosphate ions from ATP. The ATPase activity of HpSecAN68 is measured by determining the change in light absorption value to reflect the rate of NADH consumption in the enzymatic reaction using pyruvate kinase (PK) and lactate dehydrogenase (LDH) as coenzymes.
Inhibitor screening and antibacterial testing
The detailed methods are shown in Supplementary Information.
Results
Characterization and optimization of enzymatic properties of HpSecAN68
The enzymatic properties of recombinant HpSecAN68
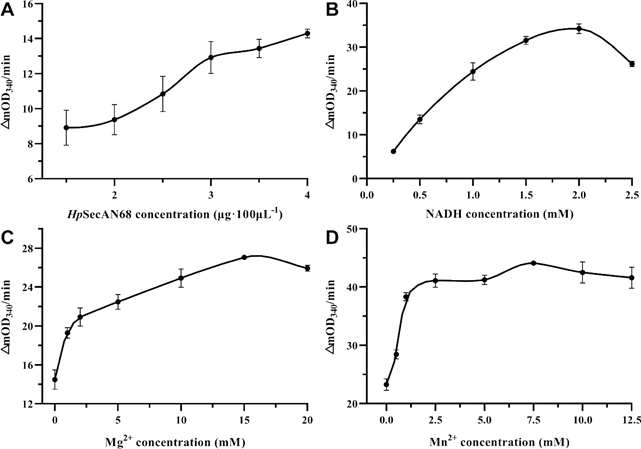
Firstly the reaction rate at different HpSecAN68 concentrations was measured. As shown in Figure 1A, the reaction rate increased with the increase in enzyme concentration, indicating that the purified protein was enzymatically active. To ensure the sensitivity of the screening, the concentration of HpSecAN68 in this reaction system was set to 3 μg/100 μL. The intermediate reactant NADH in this experiment has specific light absorption at 340 nm, and its consumption rate reflects the ATPase activity of HpSecAN68 [19]. As shown in Figure 1B, the reaction rate increases gradually with the increase of NADH concentration, reaching the maximum value when the NADH concentration is 2 mM. Divalent metal ions as the catalytic group of the enzyme active center, participate in the electron transfer and promote the binding of the substrate to the enzyme [20]. The metal ions also play important roles in the stabilization of the conformation of the enzyme [21]. As shown in Figures 1C and 1D, Mg2+, and Mn2+ have stimulatory effects on the ATPase activity of HpSecAN68. The reaction rate reached the maximum when the concentration of Mg2+ reached 15 mM, and Mn2+ reached 7.5 mM. The optimized system for the ATPase assay contains 3 μg/100 μL, 2 mM NADH, 15 mM Mg2+ and 7.5 mM Mn2+ respectively, and 12.5 mM ATP as shown below.
Determination of kinetic parameters of HpSecAN68
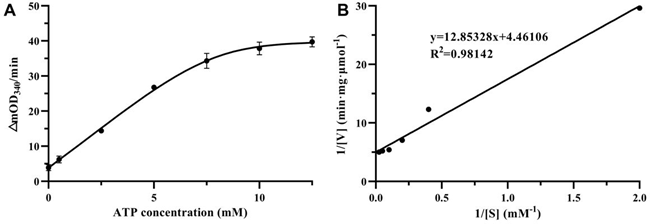
HpSecAN68 is an ATPase with ATP as its substrate, and the effect of different ATP concentrations on the conversion rate is shown in Figure 2A. The reaction rate increased gradually with the increase of ATP substrate concentration and tended to flatten when the ATP concentration reached 12.5 mM. In Figure 2B, the Lineweaver-Burk double inverse plotting method was used to plot 1/[V] versus 1/[S] to determine the kinetics of the ATPase. Under the reaction conditions of 2 mM NADH, 15 mM Mg2+, and 7.5 mM Mn2+, the Km value for HpSecAN68 to ATP was measured as 2.997 ± 0.697 μM, while the Vmax value was determined to be 0.231 ± 0.038 μmol·mg-1·min-1.
Inhibitor effects on HpSecAN68 ATPase activity
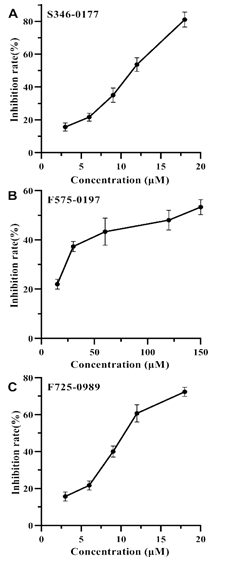
In our previous study by virtual screening [18], we identified 6 potential inhibitors against SecA, which could be classified into 3 classes based on their structural similarity. To test the inhibitory effects on HpSecAN68 ATPase activity, we selected 3 small molecule compounds from each class named S346-0177, F575-0197, and F725-0989 for verification. As shown in Figure 3, the inhibitory effects of the three compounds increased as their concentrations increased. The IC50 values calculated by fitting according to GraphPad Prism 8 software were 10.55 μM for S346-0177, 9.69 μM for F725- 0989, and 97.77 μM for F575-0197, respectively. However, it was found that when the inhibitor concentration continued to increase, all three inhibitor compounds precipitated in 5% DMSO final, which was due to the limitation of poor water solubility of the screened compounds [5, 17, 22], therefore, none of the three inhibitors could exhibit 100% inhibition effect. Nevertheless, all three compounds showed different
degrees of inhibition on HpSecAN68 ATPase activity and the inhibition showed a good dose dependence in a certain concentration range. These results show that the screened compounds indeed possess inhibiting SecA ATPase activity.
Inhibition of the growth of different types of bacteria
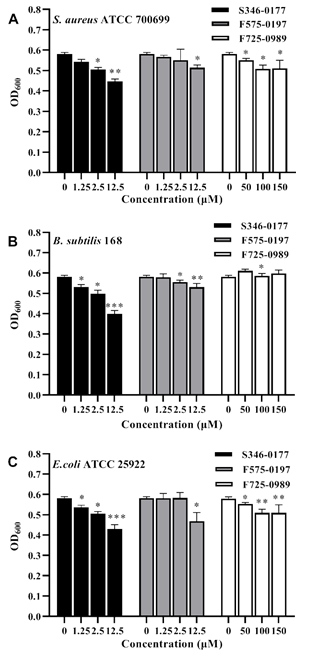
Most SecA inhibitors reported so far are not effective against Gram-negative bacteria due to the impermeability of the outer membrane [23]. So we first tested Gram-positive bacteria with single membrane such that there is no outer membrane permeability concern. Two laboratory Gram- positive bacteria, B.subtilis 168, and an important pathogen aureus ATCC 700699, were employed to validate the antibacterial activity of the three small molecules. Bacterial inhibition was expressed by 24 h growth with OD600. As shown in Figures 4A and 4B, S346-0177 from 1.25-12.5 μM and F575-0197 at 12.5 μM both have a significant inhibitory effect on the growth of B.subtilis 168 and S. aureus ATCC 700699. However, for B. subtilis 168, F725-0989 higher than 100 μM has an insignificant effect. Accordingly, we calculated the inhibition rate of each inhibitor on the growth inhibition of these bacteria. On the other hand, it has prominent antibacterial activity at 50-150 μM (P<0.05) in S. aureus ATCC 700699. The three inhibitors showed significant growth inhibition in both Gram-positive bacteria. In contrast, S346-0177 had the most significant growth inhibitory effect, and F725-0989 had a higher inhibitory concentration than the other two inhibitors. It is possible that this is related to the molecular structure and permeability of the inhibitors in the cells. E. coli SecA is the closest relative to H. pylori SecA and both are Gram-negative bacteria. Based on the above results and the high degree of conservation characterizing SecA proteins, therefore, we used E. coli ATCC 25922 to test the antibacterial activity of these inhibitors. The results are shown in Figure 4C, S346-0177 at 1.25-12.5 μM, F725-0989 at 50-150 μM were effective against E. coli ATCC 25922 in 24 h growth
incubation. F575-0197 at 12.5 μM significantly inhibited the growth of E. coli ATCC 25922, and the inhibition effect of S346-0177 was better. Interestingly, here we found that F725-0989, which had the best inhibitory effect on enzyme activity (Fig.3C), had a less significant inhibitory effect. It is not clear whether this is related to membrane permeability. We used GraphPad Prism 8 software to estimate the IC50 of the three inhibitors for three bacteria. For S. aureus ATCC 700699, the IC50 was 60.54 μM for S346-0177, 108 μM for F575-0197, and 347 μM for F725-0989. For B. subtilis 168, the IC50 were 21.16 μM for S346-0177, 57.49 μM for F575- 0197, and 208.6 μM for F725-0989. Additionally, for the Gram-negative bacterium E. coli ATCC 25922, the IC50 was 26.75 μM for S346-0177, 39.64 μM for F575-0197, and 261.1 μM for F725-0989. Regardless of the various differences, these results show that the identified small molecules have significant growth inhibition of bacteria, including the tested Gram-negative E. coli.
Discussion
pylori infection remains pretty high level in the whole world including developed and developing countries. Currently, it is difficult to eradicate H. pylori infection clinically due to the high resistance of H. pylori to many existing antibiotics. Due to the uniqueness of SecA in bacteria, SecA can be used as a novel and less toxic target for screening inhibitors against H. pylori infection and further development of new antimicrobial agents. The secretory virulence factor of H. pylori is related to the translocation of the Sec system, thus a good target for inhibitors.
The malachite green colorimetry is often used to determine free phosphate in our earlier assays for SecA ATPase [17, 24], This allows the quantitative analysis of the phosphate released by ATPase hydrolysis of ATP. To better evaluate HpSecAN68 ATPase activity and function, the NADH colorimetric method was used to determine the HpSecAN68 activity, compared with the Malachite Green Colorimetry used in the past [17, 24, 25]. HpSecAN68 was selected as a target to test its inhibitors for treating H. pylori in this study. The enzymatic kinetic parameters of HpSecAN68 ATPase were examined, and the inhibitory activities of three small molecule inhibitors against HpSecAN68 were determined. All three compounds showed significant inhibitory effects
Figure 4: Inhibitory effect of inhibitors on three species of bacteria.
(A) S.aureus ATCC 700699, (B) B. subtilis 168, (C) E.coli ATCC 25922. The data results are expressed as mean±S.D.(n=5), the on H. pylori SecAN68 ATPase activity with IC50 ranging p-values of <0.05, <0.01, and <0.001 are considered significant and are indicated with *, **, and ***, respectively. from 10.55 μM (F725-0989) to 97.77 μM (F575-0197).
Moreover, these three compounds also show inhibition of bacterial growth. Interestingly, the growth inhibition is not directly correlate with the inhibition of HpSecAN68 enzyme activity. F725-0989 showed a very significant inhibitory effect at low concentrations in the enzyme activity assay. However, in the bacteriostatic growth experiments, the inhibitory concentrations of the three bacteria were higher.
This may be due to the membrane permeability for these compounds. More interesting and perhaps more importantly, most SecA inhibitors so far cannot overcome the outer membrane impermeability of Gram-negative bacteria [23]; their inhibitory effects on Gram-negative bacteria can only be demonstrated by the use of an outer membrane mutant or chemical probe [5, 23].
We are verifying the outer-membrane permeability issue of these new identified inhibitors [23]. The current finding that these inhibitors are also active against E. coli is extremely important for further development of inhibitors against clinically important Gram-negative pathogens like H. pylori and P. aeruginosa. For laboratory safety consideration for work in progress, we are testing clinical H. pylori and some other pathogenic bacteria. The additional importance of using SecA inhibitors is that the inhibition of SecA could bypass the clinically problems of multiple-drug resistance in both Gram-positive and Gram-negative bacteria [5, 26, 27]. The water solubility of all three identified SecA inhibitors was poor, which could have some impact on the effectiveness of these compounds. Thus solving the solubility problem would be one key to the subsequent development to optimize SecA inhibitors. In further improvement of the compounds, both solubility and membrane permeability would be the critical parameters. The current identified inhibitors could be the starting lead compounds for further development of effective SecA inhibitors for antimicrobial agents. The present observation that these compounds are also effective against Gram-negative bacteria provides an additional advantage in developing general antimicrobial agents.
Conflicts of Interest
There are no conflicts of interest to declare
Acknowledgements
This work was supported in parts by the Natural Science Foundation of Sichuan Province (No. 23NSFSC0920). We would like to thank Analysis and Testing Center of Southwest Jiaotong University for providing help in equipments and analysis.
References
- Eusebi LH, Zagari RM, Bazzoli Epidemiology of Helicobacter pylori Infection. Helicobacter 19 (2015): 1-5.
- Shatalin K, Nuthanakanti A, Kaushik A, Shishov D, Peselis A, Shamovsky I, et Inhibitors of bacterial H2S biogenesis targeting antibiotic resistance and tolerance. Science 372 (2021): 1169-+.
- Economou A, Wickner W. SecA promotes preprotein translocation by undergoing ATP-driven cycles of membrane insertion and Cell 78 (1994): 835-843.
- Zimmer J, Nam YS, Rapoport Structure of a complex of the ATPase SecA and the protein-translocation channel. Nature 455 (2008): 936-U932.
- Jin JS, Hsieh YH, Chaudhary AS, Cui JM, Houghton JE, Sui SF, et SecA inhibitors as potential antimicrobial agents: differential actions on SecA-only and SecA-SecYEG protein-conducting channels. FEMS MICROBIOLOGY LETTERS 365 (2018).
- Findik BT, Smith VF, Randall LL. Penetration into membrane of amino-terminal region of SecA when associated with SecYEG in active complexes. Protein Science 27 (2018): 681-691.
- Chen L. and Tai, P.C. Evidence for ATP involvement in co-translational protein translocation. Nature, 328, 164-166 (1987). PMID: 3299104
- Jilaveanu LB, Oliver D. SecA dimer cross-linked at its subunit interface is functional for protein translocation. Journal of Bacteriology 188 (2006): 335-338.
- You, Liao M, Zhang H, Yang H, Pan X, Houghton JE, et al. Phospholipids induce conformational changes of SecA to form membrane-specific domains: AFM structures and implication on protein-conducting channels. PLoS One 8 (2013): e72560.
- Hsieh, Y-H, Zhang, H., Wang, H., Yang, H., Jiang, C., Sui S.F., and Tai, P. C. Reconstitution of functionally efficient SecA-dependent protein-conducting channels: Transformation of low-affinity SecA-liposome channels to high-affinity SecA-SecYEG- SecDF•YajC channels Biochem. Res. Commu. 431, 388-392 (2013).
- Hsieh YH, Zhang H, Lin BR, Cui N, Na B, Yang H, et SecA alone can promote protein translocation and ion channel activity: SecYEG increases efficiency and signal peptide specificity. J Biol Chem 286 (2011): 44702-44709.
- Catipovic MA, Bauer BW, Loparo JJ, Rapoport TA. Protein translocation by the SecA ATPase occurs by a power-stroke mechanism. Embo Journal 38 (2019): 13.
- Kim SH, Park M, Woo H, Tharmalingam N, Lee G, Rhee KJ, et al. Inhibitory effects of anthocyanins on secretion of Helicobacter pylori CagA and VacA toxins. Int J Med Sci 9 (2012): 838-842.
- Zawilak-Pawlik A, Zarzecka U, Zyla-Uklejewicz D, Lach J, Strapagiel D, Tegtmeyer N, et al. Establishment of serine protease htrA mutants in Helicobacter pylori is associated with secA mutations. Scientific Reports 9 (2019): 11794.
- Woo HJ, Yang JY, Lee MH, Kim HW, Kwon HJ, Park M, et al. Inhibitory Effects of β-Caryophyllene on Helicobacter pylori Infection In Vitro and In Vivo. Int J Mol Sci 21 (2020).
- Kim SH, Woo H, Park M, Rhee KJ, Moon C, Lee D, et al. Cyanidin 3-O-glucoside reduces Helicobacter pylori VacA-induced cell death of gastric KATO III cells through inhibition of the SecA pathway. Int J Med Sci 11 (2014): 742-747.
- Huang, J.; Wang, H.; Gao, F.B.; Li, M.; Yang, H.; Wang, B.; Tai, P.C. Fluorescein analogues inhibit SecA ATPase: the first sub-micromolar inhibitor of bacterial protein translocation. ChemMedChem. 2012, 7, 571-577. doi:10.1002/cmdc.201100594.
- Jian T, Su Q, Liu Y, Seoh HK, Houghton JE, Tai PC, et Structure-Based Virtual Screening of Helicobacter pylori SecA Inhibitors. IEEE Trans Nanobioscience 22, (2023) 933-942 . doi:10.1109/tnb.2023.3259946.
- Brandt M, Gale J, Anderson K, Mandichak T, Rocque W, Bickett D, et al. Expression, purification and kinetic characterization of two human acetyl-CoA carboxylase isoforms. Febs Journal 237 (2006): 369-369.
- Jiang H, Kong FD, Liu Effect of Temperature, pH and Different Metal Ions on the Activity of Digestive Enzyme from Dandong Penaeus. The Food Industry 37 (2016): 39-42.
- Zhang Y, Guo M, Zhang CC, Chen K. Study on Optimization of Chitosanase Activity of Bacillus Guangzhou Chemical Industry 47 (2019): 91-93.
- Hu JH, Akula N, Wang N. Development of a Microemulsion Formulation for Antimicrobial SecA Inhibitors. PLOS ONE 11 (2016).
- Jin J, Hsieh YH, Cui J, Damera K, Dai C, Chaudhary AS, et al. Using Chemical Probes to Assess the Feasibility of Targeting SecA for Developing Antimicrobial Agents against Gram-Negative Bacteria. ChemMedChem 11 (2016): 2511-2521.
- Hsieh YH, Huang YJ, Zhang H, Liu Q, Lu Y, Yang H, et al. Dissecting structures and functions of SecA-only protein-conducting channels: ATPase, pore structure, ion channel activity, protein translocation, and interaction with SecYEG/SecDF center dot YajC. Plos One 12 (2017): 21.
- Zhao LL, Li QP, Wei YZ, Zhang YQ, Yu LY. The Establishment and Application of Anti-Pseudomonas aeruginosa Cell-based Screening Model Targeting to SecA. Microbiology China 35 (2008): 1926-1931.
- Jin JS, Cui JM, Chaudhary AS, Hsieh YH, Damera K, Zhang H, Yang HS, Wang BH, Tai PC. Evaluation of small molecule SecA inhibitors against methicillin- resistant Staphylococcus aureus. BIOORGANIC & MEDICINAL CHEMISTRY 23 (2015): 7061-7068.
- Jin JS, Chaudhary A, Hsieh YH, Fante B, Wang BH, Tai PC. Thiouracil SecA inhibitors: bypassing the effects of efflux pumps and attenuating virulence factor secretion in MRSA and Bacillus MEDICINAL CHEMISTRY RESEARCH 30 (2021): 1341-1347.