A Systems Approach Identifies Key Regulators of HPV-Positive Cervical Cancer
Article Information
Musalula Sinkala1*, Mildred Zulu2#, Panji Nkhoma1#, Doris Kafita1, Ephraim Zulu1, Rabecca Tembo2, Zifa Ngwira2, Victor Daka3, Sody Munsaka1
1Department of Biomedical Sciences, University of Zambia, School of Health Sciences, Lusaka, Zambia
2Department of Pathology and Microbiology, University of Zambia, School of Medicine, Lusaka, Zambia
3Copperbelt University, School of Medicine, Ndola, Zambia
*Corresponding Authors: Musalula Sinkala, Department of Biomedical Sciences, University of Zambia, School of Health Sciences, P.O. Box 50110, Nationalist Road, Lusaka, Zambia
#These authors contributed equally to this work.
Received: 04 February 2021; Accepted: 19 March 2021; Published: 24 March 2021
Citation: Musalula Sinkala, Mildred Zulu, Panji Nkhoma, Doris Kafita, Ephraim Zulu, Rabecca Tembo, Zifa Ngwira, Victor Daka, Sody Munsaka. A Systems Approach Identifies Key Regulators of HPV-Positive Cervical Cancer. Journal of Bioinformatics and Systems Biology 4 (2021): 33-49.
Share at FacebookAbstract
Cervical cancer has remained the most prevalent and lethal malignancy among women worldwide and accounted for over 250,000 deaths in 2019. Nearly ninety-five per cent of cervical cancer cases are associated with persistent infection with high-risk Human Papillomavirus (HPV), and seventy per cent of these are associated with viral integration in the host genome. HPV-infection imparts specific changes in the regulatory network of infected cancer cells that are of diagnostic, prognostic and importance. Here, we conducted a systems-level analysis of the regulatory network changes, and the associated regulatory proteins thereof, in HPV-positive cervical cancer. We applied functional pathway analysis to show that HPV-positive cancers are characterised by perturbations of numerous cellular processes, predominantly in those linked to the cell cycle, mitosis, cytokine and immune cell signalling. Using computational predictions, we revealed that HPV-positive cervical cancers are regulated by transcription factors including, SOX2, E2F, NANOG, OCT4, and MYC, which control various processes such as the renewal of cancer stem cells, and the proliferation and differentiation of tumour cells. Through the analysis of upstream regulatory kinases, we identified the mitogen-activated protein kinases; among others, MAPK1, MAPK3 and MAPK8, and the cyclin-dependent kinases; among others, CDK1, CDK2 and CDK4, as the key kinases that control the biological processes in HPV-positive cervical cancers. Taken together, we uncover a landscape of the key regulatory pathways and proteins in HPV-positive cervical cancers, all of which may provide attractive drug targets for future therapeutics.
Keywords
Cervical Cancer; Human Papilloma Virus; Bioinformatics; Regulatory Networks; Signalling Pathway Analysis; Computational Predictions
Cervical Cancer articles; Human Papilloma Virus articles; Bioinformatics articles; Regulatory Networks articles; Signalling Pathway Analysis articles; Computational Predictions articles
Cervical Cancer articles Cervical Cancer Research articles Cervical Cancer review articles Cervical Cancer PubMed articles Cervical Cancer PubMed Central articles Cervical Cancer 2023 articles Cervical Cancer 2024 articles Cervical Cancer Scopus articles Cervical Cancer impact factor journals Cervical Cancer Scopus journals Cervical Cancer PubMed journals Cervical Cancer medical journals Cervical Cancer free journals Cervical Cancer best journals Cervical Cancer top journals Cervical Cancer free medical journals Cervical Cancer famous journals Cervical Cancer Google Scholar indexed journals Human Papilloma Virus articles Human Papilloma Virus Research articles Human Papilloma Virus review articles Human Papilloma Virus PubMed articles Human Papilloma Virus PubMed Central articles Human Papilloma Virus 2023 articles Human Papilloma Virus 2024 articles Human Papilloma Virus Scopus articles Human Papilloma Virus impact factor journals Human Papilloma Virus Scopus journals Human Papilloma Virus PubMed journals Human Papilloma Virus medical journals Human Papilloma Virus free journals Human Papilloma Virus best journals Human Papilloma Virus top journals Human Papilloma Virus free medical journals Human Papilloma Virus famous journals Human Papilloma Virus Google Scholar indexed journals Bioinformatics articles Bioinformatics Research articles Bioinformatics review articles Bioinformatics PubMed articles Bioinformatics PubMed Central articles Bioinformatics 2023 articles Bioinformatics 2024 articles Bioinformatics Scopus articles Bioinformatics impact factor journals Bioinformatics Scopus journals Bioinformatics PubMed journals Bioinformatics medical journals Bioinformatics free journals Bioinformatics best journals Bioinformatics top journals Bioinformatics free medical journals Bioinformatics famous journals Bioinformatics Google Scholar indexed journals Regulatory Networks articles Regulatory Networks Research articles Regulatory Networks review articles Regulatory Networks PubMed articles Regulatory Networks PubMed Central articles Regulatory Networks 2023 articles Regulatory Networks 2024 articles Regulatory Networks Scopus articles Regulatory Networks impact factor journals Regulatory Networks Scopus journals Regulatory Networks PubMed journals Regulatory Networks medical journals Regulatory Networks free journals Regulatory Networks best journals Regulatory Networks top journals Regulatory Networks free medical journals Regulatory Networks famous journals Regulatory Networks Google Scholar indexed journals Signalling Pathway Analysis articles Signalling Pathway Analysis Research articles Signalling Pathway Analysis review articles Signalling Pathway Analysis PubMed articles Signalling Pathway Analysis PubMed Central articles Signalling Pathway Analysis 2023 articles Signalling Pathway Analysis 2024 articles Signalling Pathway Analysis Scopus articles Signalling Pathway Analysis impact factor journals Signalling Pathway Analysis Scopus journals Signalling Pathway Analysis PubMed journals Signalling Pathway Analysis medical journals Signalling Pathway Analysis free journals Signalling Pathway Analysis best journals Signalling Pathway Analysis top journals Signalling Pathway Analysis free medical journals Signalling Pathway Analysis famous journals Signalling Pathway Analysis Google Scholar indexed journals Computational Predictions articles Computational Predictions Research articles Computational Predictions review articles Computational Predictions PubMed articles Computational Predictions PubMed Central articles Computational Predictions 2023 articles Computational Predictions 2024 articles Computational Predictions Scopus articles Computational Predictions impact factor journals Computational Predictions Scopus journals Computational Predictions PubMed journals Computational Predictions medical journals Computational Predictions free journals Computational Predictions best journals Computational Predictions top journals Computational Predictions free medical journals Computational Predictions famous journals Computational Predictions Google Scholar indexed journals Gene and Genomes articles Gene and Genomes Research articles Gene and Genomes review articles Gene and Genomes PubMed articles Gene and Genomes PubMed Central articles Gene and Genomes 2023 articles Gene and Genomes 2024 articles Gene and Genomes Scopus articles Gene and Genomes impact factor journals Gene and Genomes Scopus journals Gene and Genomes PubMed journals Gene and Genomes medical journals Gene and Genomes free journals Gene and Genomes best journals Gene and Genomes top journals Gene and Genomes free medical journals Gene and Genomes famous journals Gene and Genomes Google Scholar indexed journals transcriptomic articles transcriptomic Research articles transcriptomic review articles transcriptomic PubMed articles transcriptomic PubMed Central articles transcriptomic 2023 articles transcriptomic 2024 articles transcriptomic Scopus articles transcriptomic impact factor journals transcriptomic Scopus journals transcriptomic PubMed journals transcriptomic medical journals transcriptomic free journals transcriptomic best journals transcriptomic top journals transcriptomic free medical journals transcriptomic famous journals transcriptomic Google Scholar indexed journals genomic articles genomic Research articles genomic review articles genomic PubMed articles genomic PubMed Central articles genomic 2023 articles genomic 2024 articles genomic Scopus articles genomic impact factor journals genomic Scopus journals genomic PubMed journals genomic medical journals genomic free journals genomic best journals genomic top journals genomic free medical journals genomic famous journals genomic Google Scholar indexed journals proteomic articles proteomic Research articles proteomic review articles proteomic PubMed articles proteomic PubMed Central articles proteomic 2023 articles proteomic 2024 articles proteomic Scopus articles proteomic impact factor journals proteomic Scopus journals proteomic PubMed journals proteomic medical journals proteomic free journals proteomic best journals proteomic top journals proteomic free medical journals proteomic famous journals proteomic Google Scholar indexed journals
Article Details
Abbreviations:
TCGA-The Cancer Genome Atlas; GO-Gene Ontology; CB-combined score; KEGG-Kyoto Encyclopaedia of Gene and Genomes; KEA-Kinase Enrichment Analysis; ChEA-Chromatin Immunoprecipitation Enrichment Analysis; X2K-Expression2Kinases; HPV-Human Papillomavirus
1. Introduction
Cervical cancer affects more than 500,000 women and causes more than 300,000 deaths per year in women worldwide [1, 2]. Nearly ninety per cent of cervical cancer cases occur in less developed countries, where the burden of disease is disproportionately high. The disease ranks as the number one cause of female cancer mortality accounting for about a tenth of all deaths in women due to cancer [1, 2]. Cervical cancer is associated with persistent infection of high-risk human papillomaviruses (HPV). Identifying cervical cancer as having a viral aetiology has substantive consequences both for its treatment and for its prevention [1]. It is anticipated that over ninety-five per cent of cervical cancer cases will be prevented in the future by eliminating persistent infections with more than 15 oncogenic or high-risk genotypes of HPV. However, currently, cervical cancer remains mostly incurable [3]. As with other cancer-associated viruses, persistent infection of high-risk HPV genotypes is necessary for carcinogenesis of cervical cancer [4-6]. Recently, efforts have revealed that HPV virus oncogenes inactivate the tumour suppressor proteins p53 and pRB, leading to increased genomic instability, and among other cancer hallmarks, and in some cases, integration of HPV into the host genome [6, 7]. The molecular characterisation of cervical cancer by the Cancer Genome Atlas project (TCGA) and others, has provided us with a multi-dimensional understanding of the transcriptomic, genomic, proteomic, and epigenetic landscape of the disease [8-11]. These recent efforts have led to, for example, the identification of transcriptional changes of the molecularly distinct disease subtypes, the mutational landscape of cervical cancer [4, 9, 10], and the drug response and specific genetic vulnerabilities of the cervical cancer cells [12].
Despite progress made, we have barely begun to understand how some of these alterations could be leveraged to achieve effective therapeutics, owing to that we lack a multiscale understanding of how the regulatory networks of these cancers are rewired in the presence of HPV-infection. Furthermore, we are yet to identify the dysregulated pathways and the key regulators thereof that are of clinical relevance for HPV-associated cervical cancers. Here, therefore, we present a systems-level analysis of invasive cervical cancer with a focus on identifying novel alterations in the regulatory network of HPV-positive cervical cancer, the key transcription factors and kinases that may drive tumorigenesis, and thus which may all serve as markers for future therapeutic strategies.
2. Methods
We obtained uniformly processed mRNA expression data of 25 non-cancerous cervical tissue samples and 294 cervical cancer samples by the TCGA from the ARCHS4 (all RNA-seq and ChIP-seq sample and signature search) database [13, 14]. These data included mRNA transcription levels of cervical cancer samples and comprehensive de-identified clinical and sample information. The clinical information of the samples is summarised in supplementary file 1.
2.1 Identification of differentially expressed mRNA transcripts
First, we removed the genes with transcript levels less than 1 Fragment Per Kilobase Million across all samples from mRNA transcription data. Then we applied the moderated student t-test based on the negative binomial model [15, 16] to identify differentially expressed mRNA transcripts between 1) HPV-positive cervical tumours and non-cancerous cervical tissues and 2) HPV-negative cervical cancer tumours and non-cancerous cervical tissues (see Supplementary File 2). Furthermore, we defined up-regulated and down-regulated mRNA transcripts for both comparisons as those with the Benjamin-Hochberg corrected p-values [17] less than 0.05 and log2 fold-change either > 2 for the up-regulated transcripts or <-2 for the downregulated transcripts.
2.2 Pathway and functional enrichment analyses
We assessed enrichment for various Reactome pathways [18, 19] in cervical cancers by querying Enrichr [20, 21] with three gene lists for those: 1) that we found up-regulated only in the HPV-positive tumours, 2) up-regulated only in the HPV-negative tumours and 3) up-regulated in both the HPV-positive and HPV-negative tumours (see Supplementary File 3). Furthermore, we used the list of genes that we found up-regulated only in HPV-positive cervical tumours to evaluate the KEGG pathways [22], GO-term Molecular Function and GO-term Biological Processes [23] that are enriched for in HPV-positive cervical samples. We used a custom MATLAB script to connect the enriched KEGG pathway terms, GO-term Molecular Function and GO-term Biological Processes into an integrated regulatory enrichment map. Then we used software yEd to visualise the integrated enrichment landscape of the HPV-positive cervical tumours.
2.3 Prediction of the regulatory transcription factors and kinases
We identified the transcription factors and kinases that regulate the oncogenic behaviour of HPV-positive cervical cancer using a computational approach that is implemented in the software Expression2 Kinases (X2K) [24]. X2K employs a network-based reverse engineering approach to make the prediction based on prior-knowledge network data. Here, we used a list of genes that are significantly up-regulated only in the HPV-positive tumours compared to the HPV-negative tumours. Then we used the Chromatin Immunoprecipitation (ChIP) Enrichment Analysis (ChEA; 2016) [25] module of X2K to predict the upstream regulatory transcription factors that likely lead to observed transcription signature of the HPV-positive tumours (see Supplementary File 4). Next, we constructed a protein-protein interaction subnetwork by connecting the top-10 predicted transcription factors to their upstream regulators. Here, we used prior knowledge interactions from the Biological General Repository for Interaction Datasets [26], Kinase enrichment analysis database [27], KEGG pathways [28] and the Human Interaction database. Finally, we applied Kinase Enrichment Analysis to analyse the protein-protein interaction sub-network for protein kinases which are enriched for based on the number of targets that each kinase phosphorylates within the sub-network (KEA; 2015) [27]. See supplementary file 4 for a full list of the predicted kinases and their rankings based on the combined score.
2.4 Statistical Analyses
All the analyses that we have presented here were performed in MATLAB version 2020a. We tested associations between categorical variables using Fisher's exact test. Statistical significance was considered when p values were <0.05 for single comparisons, or the q-values (Benjamini-Hochberg adjusted p-values) were <0.05 for multiple comparisons.
3. Results
3.1 Clinical characteristics HPV-positive cervical cancer
We assembled a TCGA cervical cancer dataset comprising clinical information for 180 patients together with their associated mRNA transcription data. We segregated the cervical cancer samples into two groups; those that were reported HPV-positive (156 samples) and those that were HPV-negative (24 samples). We found no differences in disease outcome after the first course of treatment between HPV-positive and HPV negative cervical cancer patients (Figure1a, also see Figure S1a). Here, we also found that the reported vital statistics were not statistically different. Eighty-one per cent and eighty-three per cent of the patients were alive at the end of the follow-up period, for the patients who had HPV-positive and HPV-negative cervical cancer, respectively (Figure 1b).
The number of HPV subtypes included HPV-16 (58% of the samples), HPV-18 (18%), HPV-45 (7%), other subtypes (< 17%) (Figure S1a). HPV-45 and HPV-18 showed the highest level of integration into the host genome (100% for both virus subtypes), followed by other HPV-subtypes (92% integration) and the least level of integration into the host genome was seen among the HPV-16 subtype (76% integration) (Figure 1c). We found that 29% of cervical cancer patients older than 50 years had tumours that were associated with other HPV subtypes compared to the other age categories (< 30 years, 30 to 40 years and 40 to 50 years; 7% of samples), with a statistically significant difference, χ2 = 4.0, p = 0.045 (Figure 1d). We found no other statistically significant association between the HPV subtypes and the different age categories of cervical cancer patients.
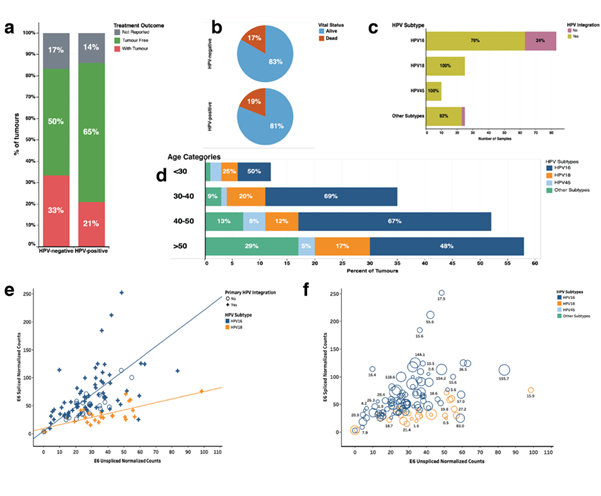
Figure 1: (a) Comparison of the clinical outcomes after the first course of treatment across the HPV-positive and HPV-negative cancer patients; (b) Pie chart showing the vital statistics after the first course of treatment across the HPV-positive and HPV-negative cancer patients; (c) Frequency of HPV subtypes in the cancer samples. The colours show the split for samples with HPV viral DNA that is integrated into the host genome; (d) Distribution of HPV-subtypes of different aged categories of tumours of patients who have HPV-positive cervical cancer; (e) Pearson's linear correlation between the normalised E6 spliced mRNA transcripts and the normalised E6 unspliced mRNA transcripts measured in HPV-positive cancers. The regression lines are plotted separately for HPV-subtypes 18 and 16. The marker shape shows labelled based on the primary HPV viral DNA integration into the host genome; (f) E6 unspliced normalised Counts E6 versus E6 spliced normalised counts. The colour shows details about HPV-subtype. The marks size shows details of the duration of overall survival (in months) of the afflicted patients. The marks are labelled with the duration of the overall survival.
We assessed the relationship between E6-spliced and E6-unspliced mRNA transcripts in the cervical cancer samples to find a significant positive correlation (r = 0.16, p<0.0001; Figure 1e). Here, we found that the Pearson linear correlation coefficient for E6-spliced verse E6-unspliced was higher for tumours infected with the HPV-16 subtype (r = 0.59, p<0.0001) compared to tumours infected with HPV-18 subtype (r = 0.29, p<0.0001). Furthermore, we found no significant association between the levels of E6 spliced and unspliced mRNA transcript and the survival outcomes of the patients who had cervical cancer (Figure 1f and Figure S1b). All the clinical information is presented in supplementary file 1.
3.2 Up-regulated genes in HPV-positive cervical cancer
We used the moderated t-test and the empirical Bayes methods to identify the differentially expressed genes in either HPV-positive or HPV-negative cervical cancers compared to the normal, non-cancerous samples. Here, we found 2,033 mRNA transcripts up-regulated in HPV-positive tumours. Among the most significantly upregulated genes were PTTG3P (log 2-fold-change [log2FC] = 11.7, q < 1 x 10-50), MMP12 (Log2FC = 10.6, q < 1 x 10-50), and BCAR4 (Log2FC = 9.5, q < 1 x 10-50; see Supplementary File 2 for all the differentially expressed genes). Most of the genes that we found significantly up-regulated in HPV-positive tumours have well-defined roles that are linked to neoplastic processes and response to viral infections. For instance, the overexpression of the pseudogene PTTG3P (Putative pituitary tumour-transforming gene 3-protein) is linked to carcinogenesis, tumour progression and metastasis in epithelial tissues in most malignancies including those of the cervix, oesophagus, breast, liver, bladder and lung [29-32]. MMP12 encodes the Macrophage metalloelastase which has antiviral immune activity and involvement in the malignant transformation of among others, cells of the cervix and skin [33-36].
3.3 Regulatory pathways and cellular processes of HPV-positive cervical cancer
We first set out to identify the dysregulated Reactome pathways among the cervical cancer samples by using the significantly up-regulated mRNA transcripts (q-value < 0.05 and Log2FC > 2). Here, to identify Reactome pathway enrichments in cervical tumours, we used three lists of up-regulated mRNA transcripts: 1) only those transcripts up-regulated in HPV-positive tumours, 2) transcripts up-regulated in both HPV-positive and HPV-negative tumours, and 3) transcripts up-regulated only in HPV-negative tumours (Figure 2). Using the 1,738 mRNA transcripts that we found up-regulated only in HPV-positive cervical cancers, we found that HPV-positive cancers exhibited a significant enrichment for 121 unique Reactome pathways (Supplementary File 3). Among the top-ranked Reactome pathways where those associated with the Cell Cycle processes (Combined Score [CS] = 188.4, q = 1.05 x 10-26), Mitotic G1-G1/S phases (CS = 122.7, 1.84 x 10-12), and DNA Replication (CS = 115.4, p = 4.33 x 10-11; Figure 2a). Conversely and unexpectedly, by focusing on the 1,574 mRNA transcripts that we found up-regulated only in HPV-negative cancers, we revealed that HPV-negative cancers exhibit significant enrichment for only 31 Reactome pathways. Here, most of the pathways enriched for are associated with G-protein coupled receptor signalling (Figure 2a).
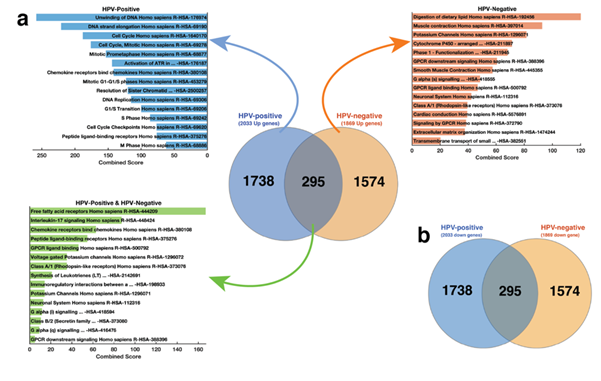
Figure 2: (a) Gene enrichment analysis plots. Top left; the top-15 Reactome pathways enriched for in HPV-positive samples. The analysis is based on the 1,738 mRNA transcripts that are up-regulated only in these samples. Centre; The distribution of significantly up-regulated mRNA transcripts among HPV-positive and HPV-negative samples. Top right; the top-15 Reactome pathways enriched for in HPV-negative samples. The analysis is based on the 1,864 mRNA transcripts that are up-regulated only in these samples; Bottom left; the top-15 Reactome pathways enriched for in both the HPV-positive and HPV-negative cervical cancers. Refer to supplementary file 3 for the complete list of Reactome pathways that represent significantly enriched genetic alterations in the HPV-positive, HPV-negative, both cervical cancers; (b) The distribution of significantly down-regulated mRNA transcripts among HPV-positive and HPV-negative samples.
Furthermore, we focused on the 295 mRNA transcripts that were up-regulated in both HPV-positive and HPV-negative tumours to reveal significant enrichment for only three Reactome pathways. These were associated with G-protein coupled ligand binding (CS = 46.4, q = 0.001), Peptide ligand-binding receptors (CS = 54.8, q = 0.005), and Rhodopsin-like receptors activity (CS = 35.4, q = 0.014; see Supplementary File 3). Surprisingly, we found that all the 2,010 mRNA transcripts that were significantly down-regulated in HPV-positive samples were also significantly down-regulated in HPV-negative samples (Figure 2b). An additional 23 mRNA transcripts were down-regulated in the HPV-negative tumours. Here, our results suggest that while the pathways whose activities are increased are distinctive among the cervical tumours that are HPV-positive and HPV-negative, those with reduced activity are comparable for both.
|
PATHWAY OR FUNCTIONAL PROCESS |
P-VALUE |
|
KEGG Pathways |
|
|
Cell cycle |
8.3 x 10-12 |
|
Cytokine-cytokine receptor interaction |
2.4 x 10-10 |
|
DNA replication |
8.5 x 10-10 |
|
Cell adhesion molecules (CAMs) |
5.9 x 10-05 |
|
Hematopoietic cell lineage |
1.4 x 10-04 |
|
GO Biological Processes |
|
|
Cytokine-mediated signalling pathway (GO:0019221) |
9.3 x 10-16 |
|
Chemokine-mediated signalling pathway (GO:0070098) |
1.0 x 10-11 |
|
The inflammatory response (GO:0006954) |
1.3 x 10-11 |
|
DNA replication (GO:0006260) |
1.7 x 10-11 |
|
Lymphocyte chemotaxis (GO:0048247) |
1.2 x 10-10 |
|
GO Molecular Function |
|
|
Chemokine activity (GO:0008009) |
4.1 x 10-12 |
|
Chemokine receptor binding (GO:0042379) |
2.0 x 10-11 |
|
Cytokine activity (GO:0005125) |
2.3 x 10-10 |
|
UDP-glycosyltransferase activity (GO:0008194) |
9.3 x 10-07 |
|
CXCR chemokine receptor binding (GO:0045236) |
3.5 x 10-06 |
Table 1: Regulatory pathways and functional processes in HPV-positive cervical cancer.
For the remaining analyses, we used the 1,738-mRNA transcript that we found up-regulated only in HPV-positive cervical tumours. Using these, first, we sought to identify the KEGG pathways, Gene Ontology (GO) terms Biology Processes and GO-term Molecular Function that are enriched for in HPV-positive cervical tumours. Here, our analyses revealed that the HPV-positive cancers are significantly enriched for 23 KEGG pathways, 175 GO-term Biological Processes and 25 GO-term Molecular Functions (Table 1, Supplementary File 3). Among the KEGG pathways, the most significantly enriched for were those associated with the Cell cycle (p = 8.33 x 10-12), Cytokine-cytokine receptor interaction (p = 2.39 x 10-10) and DNA replication (p = 8.46 x 10-10). For the GO-term Biological Processes, those that we found most significantly enriched for in HPV-positive tumours include the Cytokine-mediated signalling pathway (p = 9.34 x 10-16), Chemokine-mediated signalling pathway (p = 1.02 x 10-11), and Inflammatory response (p = 1.29 x 10-11). Furthermore, for HPV-positive tumours, the top-three significantly enriched GO-term Molecular Function were those linked to Chemokine activity (p = 4.08 x 10-12), chemokine receptor binding (p = 2.0 x 10-11), and cytokine activity (p = 2.29 x 10-10).
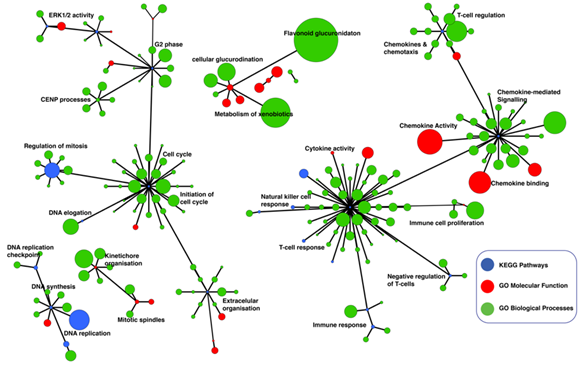
Figure 3: Network of Gene Ontology (GO) terms Molecular Functions, GO-term Biological Processes and KEGG pathways found enriched for in HPV-positive cervical cancers. Enrichr was used to obtain enriched GO-terms and KEGG pathways. The three enrichment results were combined to create an integrated network enriched functional process and pathways that were visualised in yEd (refer to the Methods section). Each node represents a GO-term Biological Process (blue), GO-term Molecular Function (red), and KEGG pathway (green). The nodes that represent similar function processes are clustered together and connected by edges. The thickness of the edges represents the number of shared genes between the nodes. The size of each node denotes the gene set size of the represented GO-term or KEGG pathway. Refer to supplementary file 3 for the complete list of KEGG pathways and GO-terms that represent significantly enriched regulatory networks in the HPV-positive cervical cancers.
We connected the significantly enriched KEGG pathway terms, GO-term Biological Processes, and GO-term Molecular Functions using known gene interactions (see Methods Sections) to reveal the overall enrichment landscape of HPV-positive cervical cancer (Figure 3). Here, we found that HPV-positive tumours are primarily driven by cytokine signalling and cell cycle processes. Overall, these findings uncover the distinct molecular mechanisms and signalling pathway perturbations which mediate oncogenic behaviour of cervical tumours that are associated with HPV infection.
3.4 Transcription factors that regulate HPV-positive cervical cancer
We used the complete list of the mRNA transcripts that we found significantly up-regulated in HPV-positive cervical cancer samples compared to the normal cervical tissues to compute enrichment for transcription factors that likely regulate the observed differential gene expression pattern. Using the in silico approach, Chromatin Immunoprecipitation Enrichment Analysis (ChEA) [25], we predicted 35 significant (p < 0.05) transcription factors that likely regulate the molecular processes that govern HPV-positive cervical cancer (Figure 4a; also see Supplementary File 4). Here, the top-ranking transcription factors were SOX2 (CS = 92.2, p = 1.51 x 10-24), E2F4 (CS = 73.3, p = 9.24 x 10-13), E2F1 (CS = 44.6, p = 1.21 x 10-10) and POU5F1 (CS = 35.0, p = 2.82 x 10-08).
Several studies show that SOX2 is overexpressed in cervical tumours and is associated with cervical neoplasia [37-39]. POU5F1 binds to the octamer motif (5'-ATTTGCAT-3') and controls the expression of several genes involved in embryonic development, oncogenesis, tumour metastasis and disease aggressiveness in several human cancers [40-42]. However, little is known about the specific roles of POU5F1 in cervical cancer. Conversely, the interplay between E6/E7 and E2F1/E2F4 in modulating the mitotic pathway, p53 pathway, pRB pathway and among other pathways is well documented [43-47].
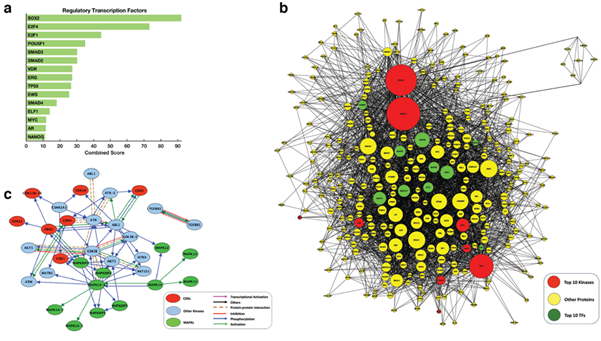
Figure 4: (a) the top-15 predicted regulatory transcription factors; (b) Intermediate protein-protein interaction subnetwork connecting transcription factors to the upstream regulatory proteins: the sub-network has 180 nodes with 1816 edges. The network is enriched for co-regulators, kinases and transcription factors that are experimentally verified to interact physically; (c) Connectivity of the top-twenty predicted kinases. Red nodes represent the cyclin-dependent kinases, the green node represent the mitogen-activated protein kinase, and cyan nodes represent other regulatory kinases. Blue nodes kinases other than the cyclin-dependent kinases or mitogen-activated protein kinases.
3.5 Regulatory kinases of HPV associated cervical cancer
We leveraged prior-knowledge protein interactions to connect the transcription factors identified using the ChEA analysis to upstream interactors and regulators to yield a protein-protein interaction sub-network (Figure 4b). The sub-network has 309 nodes that are connected by 3,299 edges. This sub-network is enriched for co-regulators, kinases and interactors of the transcription factors that drive the gene expression signature of cervical cancers infected with HPV. Next, we used Kinase Enrichment Analysis (KEA) [27] to link the proteins in the protein-protein interaction sub-network to the protein kinases that likely phosphorylate them. Here, our result was a ranked list of protein kinases that likely regulate the transcriptome signature of HPV-positive cervical tumours. Among the top-ten ranked kinases were four mitogen-activated protein kinases (MAPKs):
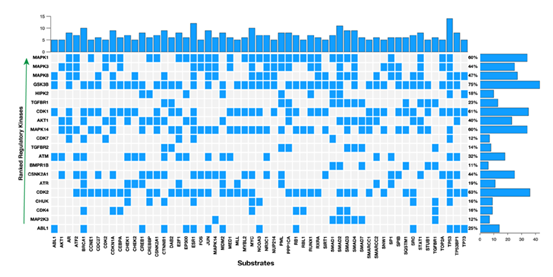
Figure 5: The top-ten predicted regulatory kinases ranked according to their combined statistical score based on the number of substrates they phosphorylate within a protein-protein interaction subnetwork. Shown along the columns of the plot are proteins which are the substrates for kinases along the rows.
MAPK1 (p = 4.82 x 10-15), MAPK3 (p = 6.56 x 10-14), MAPK8 (p = 1.71 x 10-10), and MAPK14 (p = 2.68 x 10-07; Figure 4c and Figure 5, also see Supplementary File 4). The MAPK pathway genes are altered in cervical cancers, and among other human cancers in which the MAPKs are the chief signal transduction proteins that regulate oncogenic behaviour such as cell proliferation, cell differentiation, cell survival, cancer metastasis, and resistance to drug therapy [48-52]. Accordingly, we suggest that the MAPK signalling pathways may present inflexion points for targeted therapies aimed at successfully treating HPV associated cervical cancer.
4. Discussion
We conducted a systems-level analysis to show that the biological underpinnings of HPV-positive cervical cancers are different from those of HPV-negative cancers. Our analysis revealed 121 significantly dysregulated Reactome pathways in HPV-positive cervical cancers and 31 pathways in HPV-negative cervical cancers. This finding suggests that infection with high-risk HPV genotypes additionally dysregulates several other signalling pathways, some of which may enable cancer cells to either acquire resistance or become more responsive to therapies [53-56]. We identified that the most prominent cell signalling aberrations in the HPV-positive cervical cancers were within the cell cycle and mitosis associated pathways (Figure 2a); hinting that these cancers might respond to drugs that target the cell cycle. As expected, cervical cancers have been shown to exhibit a high therapeutic response to drugs that arrest cell cycle processes [12, 57]. In this regard, we show that although alterations of multiple cancer hallmarks are required for cervical oncogenesis, those related to cell cycle processes predominantly regulate the oncogenesis in HPV-positive cervical cancer [58].
We showed that the clinical characteristics of the patients are vastly comparable for both the patients afflicted with either HPV-positive or HPV-negative cancer. Consistent with our findings, others have also shown that the overall survival outcomes of cervical cancer patients are likely not influenced by HPV infection [59]. Patients who have HPV-inactive cervical cancer have been shown to have worse outcomes compared to those who have HPV-active cervical cancer [60]. Furthermore, these findings assert the need to identify the molecular variances between cancer subtypes, which are of greater relevance to the chemotherapeutic responses of the cancers [61-65].
We integrated our findings of the altered regulatory signalling pathways (KEGG pathways), biological processes, and molecular functions to reveal the integrated regulated network of HPV-positive cervical cancers (Figure 3). Interestingly, we observed that besides alterations in the Cell cycle-related processes, HPV-positive cervical cancers show pervasive dysregulation of, among others, Cytokine-mediated processes, DNA replication, and Chemokine-mediated signalling pathway. Cytokine dysregulation in cervical cancer is a hallmark of persistent infection with high-risk HPV caused by the immunosuppressive tumour microenvironment mediated by several proinflammatory cytokines such as IL-4, IL-6, TNF-α and IFN-γ [66-69]. Strategies aimed at reversing the cytokine-related treatments are presently either being tested in clinical trials or are already in use as cervical cancer therapies with varied success [70-72].
Our computational predictions identified, among the 35 significant transcription factors, SOX2, NANOG, POU5F(OCT4), and MYC as crucial regulators of HPV-positive cervical cancer. Interestingly, the transcription factors SOX2, NONOG, MYC and OCT4 are stem cell markers that regulate the development of signalling networks including the apoptosis and cell-cycle pathways, and other pathways associated with the phenomenal proliferation and self-renewal of cells [73-75]. Since transcription factors are generally regarded as "undruggable"; we could indirectly restrain the activity of those we have identified here by inhibiting their upstream regulatory kinases [76, 77]. Concretely, we have identified several kinases of the MAP-kinase pathways, which control cell cycle-related processes including cellular proliferation, and differentiation [78-80], and kinases that directly regulate the cell cycle such as CDK1, CDK2, CDK4 and CDK7 [81, 82]. These kinases, and among others, represent credible targets for small molecule inhibitors, including CDK and MAPK inhibitors, that might be useful for controlling the disease. Notably, the pervasive alterations that we have observed in the cell cycle processes and cytokine-mediated activity provide the rationale for potentially treating HPV-positive cervical cancers using an approach that synchronously targets all these pathways.
Altogether, our analysis revealed the perturbed regulatory signalling pathways, transcription factors and protein kinases that likely govern the behaviour of HPV-positive cervical cancers, many of which are therefore potential targets for anticancer drugs.
Declarations
Author Contributions
The study was conceptualised by DK, PN, MZ, and MS; The formal methodology was devised by DK, PN, MZ, EZ, RT, VD, ZN, SM, and MS; DK, PN, RT, VD, ZN, EZ, SM, and MS performed the formal analysis of the datasets; DK, MZ, VD, RT, ZN, EZ, SM, and PN wrote the draft manuscript; the manuscript was revised by DK, PN, MZ, RT, VD, ZN, EZ, SM, and MS; DK, PN, and MS created visualisations.
Ethics Approval
The study protocol was by the University of Cape Town; Health Sciences Research Ethics Committee IRB00001938 of this study. The analyses in this study utilized publicly available datasets that were collected by the TCGA and GDSC from consenting participants. Here, all analyses were performed following the relevant policies, regulations and guidelines provided by the TCGA and GDSC for analyzing their datasets and reporting of the findings.
Competing Interests
The authors declare that they have no competing interests.
References
- Cohen PA, Jhingran A, Oaknin A, et al. Cervical cancer. Lancet 393 (2019): 169-182.
- Small W, Bacon MA, Bajaj A, et al. Cervical cancer: A global health crisis. Cancer 123 (2017): 2404-2412.
- Kessler TA. Cervical Cancer: Prevention and Early Detection. Semin Oncol Nurs 33 (2017): 172-183.
- Hu Z, Zhu D, Wang W, et al. Genome-wide profiling of HPV integration in cervical cancer identifies clustered genomic hot spots and a potential microhomology-mediated integration mechanism. Nat Genet 47 (2015): 158-163.
- Kafita D, Kaile T, Malyangu E, et al. Evidence of EBV infection in lymphomas diagnosed in Lusaka, Zambia. Pan Afr Med J 29 (2018): 181.
- Hoppe-Seyler K, Bossler F, Braun JA, et al. The HPV E6/E7 Oncogenes: Key Factors for Viral Carcinogenesis and Therapeutic Targets. Trends Microbiol 26 (2018): 158-168.
- Picksley SM, Lane DP. p53 and Rb: their cellular roles. Curr Opin Cell Biol 6 (1994): 853-858.
- Burk RD, Chen Z, Saller C, et al. Integrated genomic and molecular characterization of cervical cancer. Nature 543 (2017): 378-384.
- Huang J, Qian Z, Gong Y, et al. Comprehensive genomic variation profiling of cervical intraepithelial neoplasia and cervical cancer identifies potential targets for cervical cancer early warning. J Med Genet 56 (2019): 186-194.
- Ojesina AI, Lichtenstein L, Freeman SS, et al. Landscape of genomic alterations in cervical carcinomas. Nature 506 (2014): 371-375.
- Sinkala M, Mulder N, Patrick Martin D. Metabolic gene alterations impact the clinical aggressiveness and drug responses of 32 human cancers. Commun Biol 2 (2019).
- Zagouri F, Sergentanis TN, Chrysikos D, et al. Molecularly targeted therapies in cervical cancer. a systematic review. Gynecol Oncol 126 (2012): 29-303.
- Lachmann A, Torre D, Keenan AB, et al. Massive mining of publicly available RNA-seq data from human and mouse. Nat Commun 9 (2018): 1366.
- Weinstein JN, Collisson EA, Mills GB, et al. The Cancer Genome Atlas Pan-Cancer analysis project. Nat Publ Gr 45 (2013) 1113-1120.
- Brooks AN, Yang L, Duff MO, et al. Conservation of an RNA regulatory map between Drosophila and mammals. Genome Res 21 (2011): 193-202.
- Mortazavi A, Williams BA, McCue K, et al. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods 5 (2008): 621-628.
- Benjamini Y. Discovering the false discovery rate. J R Stat Soc Ser B (Statistical Methodol 72 (2010): 405-416.
- Fabregat A, Sidiropoulos K, Garapati P, et al. The Reactome pathway Knowledgebase. Nucleic Acids Res 44 (2016): 481-487.
- Fabregat A, Sidiropoulos K, Viteri G, et al. Reactome pathway analysis: a high-performance in-memory approach. BMC Bioinformatics 18 (2017): 142.
- Chen EY, Tan CM, Kou Y, et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinformatics 14 (2013): 128.
- Kuleshov MV, Jones MR, Rouillard AD, et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res 44 (2016): 90-97.
- Kanehisa M, Furumichi M, Tanabe M, et al. KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res 45 (2017): 353-361.
- The Gene Ontology Consortium. Gene Ontology Consortium: going forward. Nucleic Acids Res 43 (2015): 1049-1056.
- Chen EY, Xu H, Gordonov S, et al. Expression2Kinases: mRNA profiling linked to multiple upstream regulatory layers. Bioinformatics 28 (2012): 105-111.
- Lachmann A, Xu H, Krishnan J, et al. ChEA: transcription factor regulation inferred from integrating genome-wide ChIP-X experiments. Bioinformatics 26 (2010): 2438-2444.
- Oughtred R, Stark C, Breitkreutz B-J, et al. The BioGRID interaction database: 2019 update. Nucleic Acids Res 47 (2019): 529-541.
- Lachmann A, Ma’ayan A. KEA: kinase enrichment analysis. Bioinformatics 25 (2009): 684-686.
- Kanehisa M, Sato Y, Kawashima M, et al. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res 44 (2016): 457-462.
- Huang J lan, Cao S wang, Ou Q shui, et al. The long non-coding RNA PTTG3P promotes cell growth and metastasis via up-regulating PTTG1 and activating PI3K/AKT signaling in hepatocellular carcinoma. Mol Cancer 17 (2018): 93.
- Weng W, Ni S, Wang Y, et al. PTTG3P promotes gastric tumour cell proliferation and invasion and is an indicator of poor prognosis. J Cell Mol Med 21 (2017): 3360-3371.
- Lou W, Ding B, Fan W. High Expression of Pseudogene PTTG3P Indicates a Poor Prognosis in Human Breast Cancer. Mol Ther- Oncolytics 14 (2019): 15-26.
- Zhang Z, Shi Z. The pseudogene PTTG3P promotes cell migration and invasion in esophageal squamous cell carcinoma. Open Med 14 (2019): 516-522.
- Kerkelä E, Ala-Aho R, Jeskanen L, et al. Expression of human macrophage metalloelastase (MMP-12) by tumor cells in skin cancer. J Invest Dermatol 114 (2000): 1113-1119.
- Vazquez-Ortiz G, Pina-Sanchez P, Vazquez K, et al. Overexpression of cathepsin f, matrix metalloproteinases 11 and 12 in cervical cancer. BMC Cancer 5 (2005): 1-11.
- Marchant DJ, Bellac CL, Moraes TJ, et al. A new transcriptional role for matrix metalloproteinase-12 in antiviral immunity. Nat Med 20 (2014): 493-502.
- Valdivia A, Peralta R, Matute-González M, et al. Co-expression of metalloproteinases 11 and 12 in cervical scrapes cells from cervical precursor lesions. Int J Clin Exp Pathol 4 (2011): 674-682.
- Ji J, Zheng PS. Expression of Sox2 in human cervical carcinogenesis. Hum Pathol 41 (2010): 1438-1447.
- Liu XF, Yang WT, Xu R, et al. Cervical cancer cells with positive Sox2 expression exhibit the properties of cancer stem cells. PLoS One 9 (2014).
- Kim BW, Cho H, Choi CH, et al. Clinical significance of OCT4 and SOX2 protein expression in cervical cancer. BMC Cancer 15 (2015): 1015.
- Cai S, Geng S, Jin F, et al. POU5F1/Oct-4 expression in breast cancer tissue is significantly associated with non-sentinel lymph node metastasis. BMC Cancer 16 (2016): 175.
- Xu C, Xie D, Yu SC, et al. β-catenin/POU5F1/SOX2 transcription factor complex mediates IGF-I receptor signaling and predicts poor prognosis in lung adenocarcinoma. Cancer Res 73 (2013): 3181-3189.
- Takahashi K, Tanabe K, Ohnuki M, et al. Induction of Pluripotent Stem Cells from Adult Human Fibroblasts by Defined Factors. Cell 131 (2007): 861-872.
- Teissier S, Ben Khalifa Y, Mori M, et al. A New E6/P63 Pathway, Together with a Strong E7/E2F Mitotic Pathway, Modulates the Transcriptome in Cervical Cancer Cells. J Virol 81 (2007): 9368-9376.
- Rajasekaran N, Jung HS, Bae SH, et al. Effect of HPV E6/E7 siRNA with Chemotherapeutic Agents on the Regulation of TP53/E2F Dynamic Behavior for Cell Fate Decisions. Neoplasia (United States) 19 (2017): 735-749.
- Moody CA, Laimins LA. Human papillomavirus oncoproteins: Pathways to transformation. Nat Rev Cancer 10 (2010): 550-560.
- Wang W, Sima N, Kong D, et al. Selective targeting of HPV-16 E6/E7 in cervical cancer cells with a potent oncolytic adenovirus and its enhanced effect with radiotherapy in vitro and vivo. Cancer Lett 291 (2010): 67-75.
- Bester AC, Roniger M, Oren YS, et al. Nucleotide deficiency promotes genomic instability in early stages of cancer development. Cell 145 (2011): 435-446.
- Li XW, Tuergan M, Abulizi G. Expression of MAPK1 in cervical cancer and effect of MAPK1 gene silencing on epithelial-mesenchymal transition, invasion and metastasis. Asian Pac J Trop Med 8 (2015): 937-943.
- Engelbrecht AM, Gebhardt S, Louw L. Ex vivo study of MAPK profiles correlated with parameters of apoptosis during cervical carcinogenesis. Cancer Lett 235 (2006): 93-99.
- De Luca A, Maiello MR, D’Alessio A, et al. The RAS/RAF/MEK/ERK and the PI3K/AKT signalling pathways: Role in cancer pathogenesis and implications for therapeutic approaches. Expert Opin Ther Targets 16 (2012): 17-27.
- Kafita D, Nkhoma P, Zulu M, et al. Proteogenomic Analysis of Pancreatic Cancer Subtypes. BioRxiv (2020).
- Sinkala M, Nkhoma P, Mulder N, et al. Integrated molecular characterisation of the MAPK pathways in human cancers reveals pharmacologically vulnerable mutations and gene dependencies. Commun Biol 4 (2021): 1-16.
- Obenauf AC, Zou Y, Ji AL, et al. Therapy-induced tumour secretomes promote resistance and tumour progression. Nature 520 (2015): 368-372.
- Ramos P, Bentires-Alj M. Mechanism-based cancer therapy: resistance to therapy, therapy for resistance. Oncogene 34 (2015): 3617-3626.
- Pagliarini R, Shao W, Sellers WR. Oncogene addiction: pathways of therapeutic response, resistance, and road maps toward a cure. EMBO Rep 16 (2015): 280-296.
- Gazdar AF, Shigematsu H, Herz J, et al. Mutations and addiction to EGFR: The Achilles “heal” of lung cancers? Trends Mol Med 10 (2004): 481-486.
- Prasad SB, Yadav SS, Das M, et al. PI3K/AKT pathway-mediated regulation of p27Kip1 is associated with cell cycle arrest and apoptosis in cervical cancer. Cell Oncol 38 (2015): 215-225.
- Hanahan D, Weinberg RA. Hallmarks of cancer: The next generation. Cell 144 (2011): 646-674.
- Rodríguez-Carunchio L, Soveral I, Steenbergen R, et al. HPV-negative carcinoma of the uterine cervix: a distinct type of cervical cancer with poor prognosis. BJOG An Int J Obstet Gynaecol 122 (2015): 119-127.
- Banister CE, Liu C, Pirisi L, et al. Identification and characterization of HPV-independent cervical cancers. Oncotarget 8 (2017): 13375-13386.
- Samarzija I, Beard P. Hedgehog pathway regulators influence cervical cancer cell proliferation, survival and migration. Biochem Biophys Res Commun 425 (2012): 64-69.
- Koren S, Bentires-Alj M. Breast Tumor Heterogeneity: Source of Fitness, Hurdle for Therapy. Mol Cell 60 (2015): 537-546.
- Chien W, Sudo M, Ding LW, et al. Functional genome-wide screening identifies targets and pathways sensitizing pancreatic cancer cells to dasatinib. J Cancer 9 (2018): 4762-4773.
- Barretina J, Caponigro G, Stransky N, et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 483 (2012): 603-607.
- Weinstein IB, Joe A. Oncogene addiction. Cancer Res 68 (2008): 3077-3080.
- Audirac-Chalifour A, Torres-Poveda K, Bahena-Román M, et al. Cervical microbiome and cytokine profile at various stages of cervical cancer: A pilot study. PLoS One 11 (2016).
- Paradkar PH, Joshi JV, Mertia PN, et al. Role of cytokines in genesis, progression and prognosis of cervical cancer. Asian Pacific J Cancer Prev 15 (2014): 3851-3864.
- Song Z, Lin Y, Ye X, et al. Expression of IL-1α and IL-6 is associated with progression and prognosis of human cervical cancer. Med Sci Monit 22 (2016): 4475-4481.
- Berti FCB, Pereira APL, Cebinelli GCM, et al. The role of interleukin 10 in human papilloma virus infection and progression to cervical carcinoma. Cytokine Growth Factor Rev 34 (2017): 1-13.
- Peralta-Zaragoza O, Bermúdez-Morales VH, Pérez-Plasencia C, et al. Targeted treatments for cervical cancer: A review. Onco Targets Ther 5 (2012): 315-328.
- Thanendrarajan S, Nowak M, Abken H, et al. Combining cytokine-induced killer cells with vaccination in cancer immunotherapy: More than one plus one? Leuk Res 35 (2011): 1136-1142.
- Alshaker HA, Matalka KZ. IFN-γ, IL-17 and TGF-β involvement in shaping the tumor microenvironment: The significance of modulating such cytokines in treating malignant solid tumors. Cancer Cell Int 11 (2011): 33.
- Van Schaijik B, Davis PF, Wickremesekera AC, et al. Subcellular localisation of the stem cell markers OCT4, SOX2, NANOG, KLF4 and c-MYC in cancer: A review. J Clin Pathol 71 (2018): 88-91.
- Liu X, Huang J, Chen T, et al. Yamanaka factors critically regulate the developmental signaling network in mouse embryonic stem cells. Cell Res 18 (2008): 1177-1189.
- Okita K, Yamanaka S. Induction of pluripotency by defined factors. Exp Cell Res 316 (2010): 2565-2570.
- Hagenbuchner J, Ausserlechner MJ. Targeting transcription factors by small compounds - Current strategies and future implications. Biochem Pharmacol 107 (2016): 1-13.
- Masamoto Y, Kurokawa M. Targeting chronic myeloid leukemia stem cells: can transcriptional program be a druggable target for cancers? Stem Cell Investig 5 (2018).
- Sun Y, Liu WZ, Liu T, et al. Signaling pathway of MAPK/ERK in cell proliferation, differentiation, migration, senescence and apoptosis. J Recept Signal Transduct 35 (2015): 600-604.
- Keshet Y, Seger R. The MAP kinase signaling cascades: a system of hundreds of components regulates a diverse array of physiological functions. Methods Mol Biol 661 (2010): 3-38.
- Zhang Y, Dong C. Regulatory mechanisms of mitogen-activated kinase signaling. Cell Mol Life Sci 64 (2007): 2771-2789.
- Malumbres M. Cyclin-dependent kinases. Genome Biol 15 (2014): 122.
- Echalier A, Endicott JA, Noble MEM. Recent developments in cyclin-dependent kinase biochemical and structural studies. Biochim Biophys Acta - Proteins Proteomics 1804 (2010): 511-519.
Supplementary Figure 1:
(a) Kaplan-Meier 1 curves of the duration overall survival of patients with HPV-positive and HPV-negative cervical cancer; (b) normalised spliced E6 versus overall survival month. The markers are coloured based on the HPV-subtypes found the samples.
References
- Goel MK, Khanna P, Kishore J. Understanding survival analysis: Kaplan-Meier estimate. Int. J. Ayurveda Res 1 (2010): 274-278.
