Animal Model Studies Reveal that Common Human-Centric Non-Coding Variants from Epidemiology are By-products of Primate Evolution Unrelated to Physiological Control of Blood Pressure
Article Information
Alan Y Deng*, Annie Menard
Research Centre, CRCHUM (Centre hospitalier de l’Université de Montréal), Department of Medicine, Université de Montréal, Montréal, Québec, Canada
*Corresponding Author: Alan Y Deng, Research Centre, Centre hospitalier de l’Université de Montréal (CHUM), 900 rue St. Denis Street, R08-432, Montréal, Québec, H2X 0A9, Canada
Received: 23 August 2021; Accepted: 30 August 2021; Published: 15 September 2021
Citation: Alan Y Deng, Annie Menard. Animal Model Studies Reveal that Common Human-Centric Non-Coding Variants from Epidemiology are By-products of Primate Evolution Unrelated to Physiological Control of Blood Pressure. Cardiology and Cardiovascular Medicine 5 (2021): 471-501.
Share at FacebookAbstract
Background: Human genome-wide association studies (GWAS) on blood pressure (BP) have been undertaken by avoiding its physiology and mechanisms controlling BP. Consequently, the physiological significance of GWAS on BP remains undiscovered. A shared mechanistic foundation starts to untangle human physiological regulations of BP as primate versions of rodents and vice versa. Thus, understanding mechanisms in rodents is equivalent to unraveling the same in humans rooted in their common ancestors.
Methods: We used BP quantitative trait loci (QTLs) from hypertensive rats as functional proxies to seize human orthologs marked by GWAS.
Results: 6 BP QTLs correspond to 6 specific human genes. BP was altered by these QTL alleles, and yet, the human non-coding GWAS variants are absent in rodents. They cannot contribute to physiological modulations of BP by these QTLs, because depleting such a variant has no impact on BP. Thus, these variants mark QTLs nearby, are not QTLs per se, since they only emerged during primate evolution. When functioning together, these human QTLs physiologically attain the same magnitude of BP effect as a single QTL alone. Mechanistically, these QTLs may function in a common pathway. Each is involved in a different pathway step leading to BP control, not by altering BP by merely affecting QTL expressions. One pathway is muscarinic cholinergic receptor 3 (M3R) signaling. A new M3R component is implicated from current work.
Conclusions: In spite of cognitive impedance from a human-centric dogma, the modularity/pathway concept is evolving into a paradigm physiologically applicable to mammalian polygenic and quantitative traits.
Keywords
Quantitative trait loci; Modularity; Common pathway; Epistasis; GWAS
Quantitative trait loci articles; Modularity articles; Common pathway articles; Epistasis articles; GWAS articles
Article Details
1. Introduction
1.1. A high prevalence of chronically-elevated blood pressure, hypertension, is a compelling risk for cardiovascular, renal and infectious diseases [1]. This risk has been highlighted by a recent hospitalization rate of COVID-19 patients with underlying conditions (DOI: 10.15585/mmwr.mm6915e3). The most common among them is hypertension that needs to be treated with anti-hypertensive drugs. This alarming hazard urgently demands our actions in unraveling pathogeneses of hypertension, and in distinguishing mechanistic causes of pathophysiology from their outward effects found in epidemiology.
1.2. Thanks to genome-wide association studies (GWASs), detecting human quantitative trait loci (QTLs) for blood pressure (BP) has statistically marked the vicinity of more than 900 BP QTLs by more than 10000 single nucleotide polymorphisms (SNPs) [2]. So far, no human QTLs have been functionally identified to belong to a physiology system known to affect blood pressure. We are no closer in understanding a pathogenesis for human polygenic hypertension than before the advent of GWASs [3]. With due respect, >90% of these SNPs cannot be functional variants in spite of the equally-strong statistics associating them all with blood pressure. These SNPs are pure markers for potential QTLs close by. Thus, in identifying a human QTL, statistics are insufficient and physiological studies in vivo are needed.
1.3. A physiological distinction is unmistakable between locating a SNP marking a QTL nearby and identifying the QTL itself. By focusing on after effects of BP-regulating mechanisms, human GWASs have reached their limitations in finding causes of these mechanisms. A QTL refers to a locus residing in a chromosome segment when genetically defined, but a QTL is a single gene when molecularly identified [4]. For example, C17QTL1 on rat Chromosome 17 is a single gene encoding Chrm3 [muscarinic cholinergic receptor 3 (M3R)] [5,6]. No combination with another QTL/gene is necessary to physiologically affect blood pressure [7,8].
1.4. Functionally, an alteration in a physiological mechanism will cause variations in blood pressure, but not all BP variations are a result of mechanistic changes. Experimental advantages using rodent models allow causative mechanisms modulating blood pressure driven by physiology to be unveiled [9]. Because of conserved mechanisms, studying them in regulating rodent BP is equivalent to revealing the same mechanisms in humans originating from their common ancestors. Evidently, most land living mammals attain a similar range of blood pressures [10], despite differences in separate physiology characters (e.g. corporal bulk). The only way for this to materialize is that basic mechanisms modulating blood pressure must have been formed and held constant in common ancestors of these mammals before 90 million years ago (www.timetree.org), before humans existed.
1.5. In vivo studies now unify animal model and human QTLs into a basic framework in physiological mechanisms of BP control. Rodent QTLs as proxies from inbred strains have functionally captured distinct human QTLs [11,12]. The intergenic GWAS SNP close to CHRM3 [13] is only a marker for the human QTL, not the QTL itself [11]. Thus, a ‘common’ SNP from human GWAS merely marks a nearby physiological QTL that has a ‘rare’ functional variant. Replicating such a ‘common’ SNP by other GWAS seems an epidemiological exercise [2] that is irrelevant to its physiological impact on BP, because removing the SNP has no effect on BP [5,7].
1.6. These atypical and counter-intuitive results may appear confusing and disturbing, because they contradict the prevailing tenet in epidemiology that a quantitative and polygenic trait should be made by accumulating ‘miniscule’ effects from multiple QTLs. This is lately molded into an ‘omnigenic’ hypothesis [3], which has been believed to universally apply to all polygenic and quantitative traits in whatever organisms in both inbreds and oubreds. (Comparing inbreds to outbreds will be elaborated further in discussions). However, facts are facts. There is no in vivo evidence that any of human genetic architectures of GWAS in outbreds [2] could actually impact on blood pressure physiologically, and controlling gene expressions from a GWAS SNP might affect BP physiologically.
1.7. The shift of paradigm from ‘omnigenicity’ to QTL modularity [11,12,14] has become a part of the literature in polygenic research [15]. Invisible QTL modularity from human epidemiology reflects limitations of GWAS [2], because GWAS is done by ignoring BP-controlling physiology and mechanisms. Among mammals, modularity is a physiological reality [9] but hidden from GWAS. It is analogous to lacking evidence for black holes in Newton’s mechanics, although they exist from Einstein’s theory of gravity [9]. It seems that physiologically understanding quantitative and polygenic traits such as BP in biology mimics studying gravity in physics.
1.8. Only by accumulating evidence can we make this paradigm shift recognizable and accepted by the concerned scientific community. In retrospect, the inherent truth embodied in Mendelism became established and appreciated after a 35-year oblivion, only when Mendel’s discoveries were reproduced by others. Hopefully, our previous work [11,12,14] and the current confirmation of them, will encourage different scientists to expand one-lab-based findings in animal models and humans to broader polygenic traits including BP. In this way, the validity of this developing paradigm can be further tested.
1.9. Arguably, a narrow range of work [11,12] might not represent a broad mechanistic reality. We have extended the rat QTL coverage to evaluate additional human GWAS genes in progressive stages, as a lone investigator-initiated lab can. Instead of replicating same QTLs in other populations as human GWASs do [3], we tested the reproducibility of mechanistic outcomes from analyzing previously-unexplored rat QTLs that respond to different human GWAS gene orthologs. In this process, our new data have validated and widened the paradigm of QTL modularity to both rodents and humans as pathogenic pathways to polygenic hypertension [9]. Previously-unsuspected components of these pathways have been implicated.
2. Materials and Methods
2.1. Animals Protocols for handling as well as maintaining animals were approved by our institutional animal committee (CIPA). Inbred hypertensive Dahl salt-sensitive rats (DSS) are our functional proxy of choice. In order to detect the physiological impact from a BP QTL, our work was done in the DSS genetic background that has lost its genome buffering capacity in impeding BP fluctuations [16] and in suppressing hypertension [17,18].
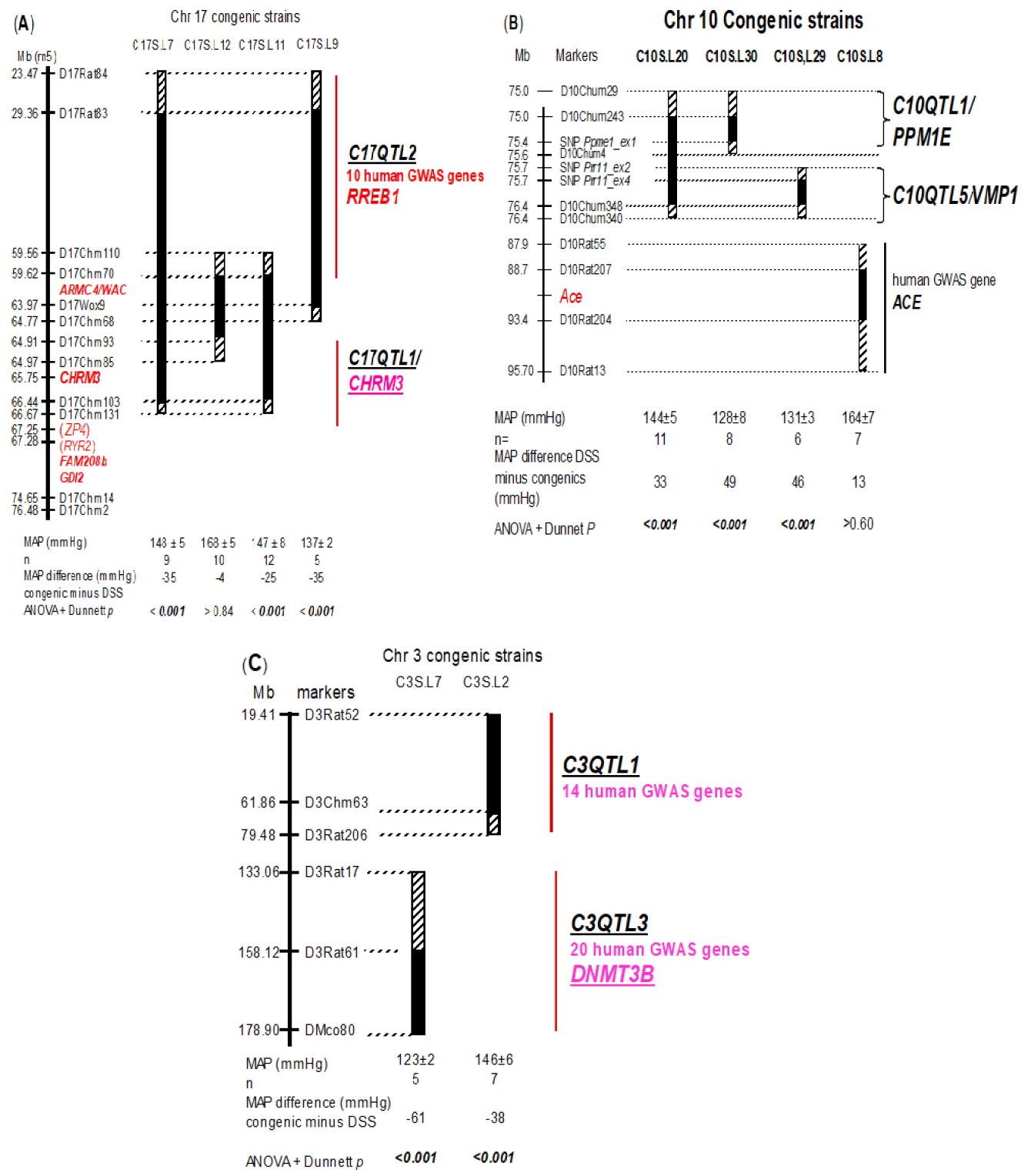
2.2. Experimental protocols and analyses Breeding procedure, dietary treatments, telemetry implantation, postoperative care and BP measurement durations were essentially the same as reported previously [14]. In brief, male rats were weaned at 21 days of age, kept on a low salt diet followed by a high salt diet starting from 35 days of age until the end of the experiment. Telemetry probes were implanted at 56 days of age (namely 3 weeks from the time of the high salt diet). In the BP presentation (Figure 1), averaged readings of mean arterial pressures (MAP) for the duration of measurement were given for each strain.
Figure 1: Congenic knock in genetics defining chromosome regions containing BP QTLs in vivo. Solid bars under congenic strains symbolize the Dahl salt-sensitive (DSS) chromosome fragments that have been replaced by those of Lewis. Hatched bars indicate ambiguous regions. The full gene names with abbreviations is given in the Table 1 legend. Mean arterial pressures (MAPs) for DSS and congenic strains are averaged for the period of measurement and are given at the bottom of the map. Significant p values are put in bold and italics. ± indicates SEM. (A) Chr 17; (B) Chr 10; (C) Chr 3.
2.3. Repeated measures’ analysis of variance (ANOVA) followed by Dunnett’s test, which corrects for multiple comparisons and unequal sample sizes, was used to compare a parameter in MAP between 2 groups as reported previously (14). The power and sample size calculations in the analysis are the same as given previously [11].
3. Results
3.1. Congenic knock in genetics is a proxy tool in physiologically catching human GWAS genes by causality. The congenic principle is similar to that of SNP ‘knock-in’ with a variation in a genome scale [4], and is employed as congenic knock in genetics [11,12]. Despite the DSS rats are known for their ‘salt-sensitivity’ for physiological studies, the human GWAS genes that have been captured by the DSS model commonly function in humans from general populations, with or without salt sensitivity [11,12]. Furthermore, the M3R signaling pathway found in DSS is pro-hypertensive even under low salt diet [5]. High salt diet merely accelerated hypertension and our studies of it using the model. Thus, BP-regulating mechanisms discovered in DSS rats are applicable to those of human essential hypertension in general populations, irrespective of the salt content. Among DSS chromosome segments known to contain BP QTLs [14], only those matching human QTL signals from GWAS [2] were investigated here (Supplementary Table 1).
3.2.1. The M3R signaling pathway in regulating BP existed in common ancestors of humans and rodents. Since they have similar blood pressures in a polygenic and quantitative context, we hypothesized that humans and rodents may use same pathways originating from their common ancestors and their similar BP states are not due to a convergent evolution event. We tested this hypothesis by focusing on M3R as C17QTL1, a QTL on rat Chromosome 17 and human CHROMOSOME 1, because M3R has been physiologically proven to be C17QTL1 [5,7,11].
3.2.2. Despite no M3R sequence is available from extinct common ancestors of humans and rodents 90 million years ago, the M3R signaling already existed in them. This is because M3R of Tasmania Devil, a marsupial, shares 90% of homology conservation with humans and rats overall, and 95% in the M3R signaling domain to either humans or rats (Supplementary Table 2, http://genome.ucsc.edu). M3R’s presence in them indicates that the M3R signaling pathway was present even in common ancestors of marsupial and placental mammals. Marsupials and placentals split 160 million years ago, prior to the divergence of rodent and human ancestors 90 million years ago, and before modern humans existed around 300 thousand years ago (www.timetree.org). Marsupials have similar BP as most placental land mammals [10].
3.2.3. This is a proof that, common ancestors of humans and rodents possessed a BP-regulating mechanism of M3R signaling pathway, in spite of the fact that M3R is pleiotropic in functions in addition to regulating BP. This evidence explains similar BPs between humans and rodents, for which a possible convergent evolution by different mechanisms to obtain similar blood pressures has no proof. This contrast will be dealt with further in discussions.
3.3.1. Defining physiological effect of each QTL/GWAS gene on blood pressure: GWAS SNPs were chosen solely by their elevated minor allele frequencies, not based on their influences on blood pressure. Total variance used in GWAS [3] gauges a spread of BP in heterogeneous populations as an epidemiology parameter, and thus is irrelevant to physiological mechanisms of BP control. Total variance is largely due to environmental effects, not due to mechanistic actions of QTLs [9]. Since environmental factors are not inherited and mostly unquantifiable, the often-termed ‘missing’ heritability is not equivalent to missing total variance. Regardless how many BP QTLs really exist in an individual organism, identifying the physiological impact of a particular QTL on BP is critical that can be untangled from the non-physiological total variance
3.3.2. The BP effect for a human GWAS SNP [2] and presumably from one QTL marked by it, seemed ‘miniscule’ when fractionated from total variance [3]. What then is the significance in identifying 6 ‘trifle’ QTLs among 900 [2], each encoded by a single gene (Table 1, Figure 1)? An illuminating answer came from revealing the actual magnitude of physiological BP effect for each of these 6 QTLs. It turns out to be substantially larger when physiologically defined, alone in homogeneity, by causality, and under a uniform environment [details were given in Table 1 in reference] [19].
Table 1: Selective rat QTLs with missense mutations functionally capturing human orthologs.
Footnote to Table: QTLs and their BP effects are given in Fig. 1. § shows confirmed mutations; * indicates the functional impact identified. Detailed data are presented in Supplemental tables. CHRM3, muscarinic cholinergic 3 receptor (M3R) gene; DNMT3B, DNA methyltransferase 3 beta; PPM1E, protein phosphatase, Mg2+/Mn2+ dependent 1E; RREB1, ras responsive element binding protein 1; VMP1, vacuole membrane protein 1.
3.3.3. Here, a distinction is revealing between a statistical fractionation from total variance and a real physiological impact for each QTL. This is because a fractionated BP effect for every one of these 900 human QTLs in GWAS [2] was compounded by those from other genes in heterogeneity, and by environmental influences in study populations. Due to these interferences, an estimated effect for each QTL from total variance does not prove if the GWAS signal itself can have a physiological impact on BP, let alone its magnitude of BP effect. C17QTL2 on DSS rat Chromosome 17 proves the point (Figure 1A).
3.3.4. Ten different human GWAS genes [2] fell into the congenic knock-in segment that defined C17QTL2 (Figure 1A). One human GWAS signal close to CHRM3 [13] was present in the segment containing C17QTL1. The 2 QTLs were detected as a single QTL statistically and explained 6.7% of total variance in a heterogeneous rat population [20]. In contrast, each of them was capable of physiologically and independently altering BP by 28-42% in the total difference by mmHg in vivo between 2 parental rat strains and under a uniform environment (Table 1). The calculation is as follows.
3.3.5. The congenic knock-in defining C17QTL2 lowered blood pressure by 35 mmHg (Figure 1A). The total BP difference between DSS and Lewis strains is 83 mmHg, i.e. 178 mmHg for DSS minus 95 mmHg for Lewis. Thus, the physiological BP effect of C17QTL2 is calculated as 35/83 = 42%. By the same calculation, each of 4 QTLs, C10QTL1, C10QTL5, C3QTL1, and C3QTL3, singularly possesses a physiological BP effect ranging from 46-76% (Figure 1, Table 1). This is contrary to the non-physiological estimate from fractionated total BP variance [2].
3.3.6. If the ‘miniscule’ hypothesis were physiologically valid [3], eliminating 1 QTL/GWAS gene among 900 should have a negligible consequence. This is not the case, since Chrm3 is a single gene responsible for C17QTL1. Depleting C17QTL1/Chrm3 alone lowered BP by at least 50% in the BP difference between Chrm3+/+ and Chrm3-/- (5). Cumulatively, the 6 QTLs by themselves (Table 1) seem physiologically more than sufficient to explain the total BP difference between the 2 parental strains.
3.3.7. Therefore, the crucial issue to address is not how ‘miniscule’ could be the BP effect that each GWAS gene was supposed to have, but rather why there is such an over-abundance of GWAS genes that are more than physiologically necessary in regulating BP of an organism. Summing them up cannot be a valid physiological justification, whereas fractionating each from total variance is not physiological. Thus, a meaningful physiological solution is required on the collectivity of their functional impact on BP.
3.4.1. QTL modularity on BP is physiologically conserved between humans and rodents: We first combined C17QTL1 and C17QTL2 in a ‘double’ congenic strain as shown in C17S.L7. Their aggregated MAP (148±5, n=9) was similar to either of them alone (Figure 1A). A 2x2 ANOVA (14) showed epistasis (p <0.003) between them, i.e. their combined BP is non-cumulative and they belong to the same epistatic module, epistatic module 2. Epistasis means one QTL hiding the effect of another and occurs regardless the number of GWAS genes involved. The C17QTL2-residing segment bears 10 human GWAS genes and the C17QTL1-residing segment carries 1 human GWAS gene (Figure 1A).
3.4.2. The congenic strain C10S.L20 (Figure 1B) contains C10QTL1 and C10QTL5 and showed similar BP (144±5, n=11) as either of the 2 QTLs alone. There is epistasis (p <0.001) between the 2 QTL-residing intervals carrying 4 human GWAS genes together (Figure 1B, Supplementary Table 1). Either of C3QTL1 and C3QTL3 showed epistasis with C10QTL1 [14]. Thus, these 4 QTLs belong to the same module, epistatic module 1. The chromosome segments lodging C3QTL1 and C3QTL3 lodge 34 human GWAS genes (Figure 1C, Supplementary Table 1).
3.4.3. In total, 49 human GWAS genes are contained in the chromosome segments harboring the 6 QTLs (Figure 1). On the basis of their known epistasis by functional proxy, these 6 human GWAS genes may be classified physiologically into 2 epistatic modules, or 2 independent pathways in determining BP. BP is functionally additive between 2 members of 2 separate modules [14], and is the basic mechanism of QTL actions. Since M3R is a signaling pathway [5], epistatic module 2 to which C17QTL1/CHRM3 and C17QTL2 belong constitutes a pathway with multiple steps composed of different QTLs leading to BP control. C17QTL2 most likely participates in one of these steps in the M3R signaling pathway.
3.4.4. In order to identify a specific step in a pathway, a molecular identification of a QTL is necessary. Identifying CHRM3 as a causal gene to C17QTL1 is the precedent for genetically discovering a component of a step in a pathway in a polygenic context [5,6]. CHRM3 has been proven to be C17QTL1 not only in DSS rats [5], but also a strong candidate for humans [11]. Chrm3 carries a function-changing missense mutation [5]. Thus, missense mutations are priority, although not exclusive, targets for identifying candidate genes for the following QTLs.
3.5.1. C17QTL2 of DSS rats may be a physiological ortholog of a human GWAS gene, RREB1 (ras responsive element binding protein 1). 10 different human GWAS genes (2) fell into the large congenic knock-in segment that defined C17QTL2 by changing BP in vivo (Figure 1A). It appears that at least 3 human QTLs among the 10 GWAS genes may exist in this interval, since they are located on 3 separate human CHROMOSOMEs (CHRs) (Supplementary Table 1). Of 6 GWAS genes on CHR 6, RREB1 has become a functional candidate gene for C17QTL2, because it carries 2 missense mutations in DSS rats that may potentially alter the function of the Rreb1 protein (Supplementary Table 3).
3.5.2. The human RREB1 gene was marked by 2 intronic SNPs and 1 missense mutation (Supplementary Tables 1&3). If any of them would affect BP in vivo, it should be present in rodents. However, no similar SNP sequence, nor homology in 1 kb non-coding sequence surrounding the 2 intronic SNPs, were detected in the rat genome (Table 2). These non-coding SNPs are present only among primates including humans (Table 3). Since BPs of rodents and primates are similar (10), these non-coding SNPs seem a by-product of primate evolution, rather than a requirement in physiologically controlling BP. Since BP changed in vivo without them (Fig. 1A), these SNPs themselves cannot be responsible for changing BP by C17QTL2, and appears solely as human-centered markers for the physiological QTL nearby.
|
Human SNP/ Marked gene |
Rat Homology |
Note |
|
rs4960295/RREB1 (intron) |
No |
No hits in 1Kb sequence used for blast on Chr17 |
|
rs2151942/RREB1 (intron) |
No |
Haphazard hits in 1 Kb sequence used for blast; 2 mini regions of homology randomly distributed, but not in the right region. |
|
rs1334576/RREB1 (missense Gly195Arg) |
SNP is absent |
91 bases hit (93.4% homology) in 1 Kb sequence used for blast on Chr17 |
|
rs6141767/DNMT3B (intron) |
No |
Similar to rs2151942/RREB1 |
|
rs304295/PPM1E (intergenic) |
No |
Similar to above |
|
rs304298/PPM1E (beginning of intron1) |
No |
Similar to above |
|
rs12942969/PPM1E (middle of intron1) |
No |
No hits in 0.6Kb sequence used for blast on Chr10 |
|
rs35082135/PPM1E (end of intron1) |
No |
No hits in 1Kb sequence used for blast on Chr10 |
|
rs2645466/VMP1 (intron3) |
No |
Similar to rs2151942/RREB1 |
|
rs2820037/CHRM3 (intergenic) |
No |
No hits in 4Kb sequence used for blast on Chr17 |
Table 2: A survey of sequence homologies between humans and the rat for GWAS SNPs
Footnote: Gene names are given in legends of Table 1. Appropriate sequence surrounding a SNP in question was blasted into the rat genome at: https://genome.ucsc.edu/cgi-bin/hgGateway.
More recent more ancient
Table 3: Survey of conservation/homology for non-coding GWAS SNPs and 1 missense mutation during primate evolution
Footnote: Gene names are given in the legend for Table 1. Homology indicates that the SNP and/or surrounding sequences are conserved. Old World monkeys are represented by Rhesus macaque, baboon; New World monkeys are represented by marmoset, squirrel monkey. Searches were done at https://genome.ucsc.edu/cgi-bin/hgGateway.
3.5.3. The physiological C17QTL2 has to be both conserved between the rat and human as well as capable of potentially altering BP by function. The coding domain of f RREB1 fulfils these 2 criteria. First, missense mutations modifying its protein structure may have a function impact. They are the human rs1334576 missense mutation changing Gly195Arg, and 2 missense mutations in DSS rats (Supplementary Table 3). Additional missense mutations [21] are found in the human RREB1 (Table 4). In contrast to the 2 intronic SNPs (Table 2), RREB1 coding regions are highly conserved between rodents and humans. Although the coding mutations in humans are not the same as in inbred DSS rats, they may affect the function of RREB1 from different positions of amino acids. This shared feature and presence of coding mutations support the candidacy of the RREB1 protein for C17QTL2 for both humans and DSS rats.
Table 4: Amino acid alignment and missense mutations in codons for ras responsive element binding protein 1 (RREB1) in humans and rats
Footnote for table: * indicates amino acid identity (86%). Probable human missense mutations, (21) are shaded. Amino acids in blue indicate that the minor allele has been observed more than 10 times. The human Gly195Arg is a GWAS coding SNP. The 2 rat missense mutations are detected in both our database, (36) and rat genome database, (37) and shaded in green. DSS, Dahl salt-sensitive rats.
3.6.1. C3QTL3 may be a physiological ortholog of human GWAS gene DNA methyltransferase 3B (DNMT3B). The knock-in segment defining C3QTL3 is large, and contains 20 human GWAS genes located on human CHR 20 (Supplementary Table 1). Among them, 5 rat orthologs for 5 GWAS genes carry synonymous mutations. 1 rat ortholog for DNMT3B carries a missense mutation changing Met80Val (Supplementary Tables 3&4). This function-altering potential made the DNMT3B protein a physiological candidate for C3QTL3 conserved in basic mechanisms of BP physiology between rats and humans. The human DNMT3B coding region carries several missense mutations (Supplementary Table 4).
3.6.2. In contrast, the intronic human GWAS SNP, rs6141767, marked DNMT3B, is absent in the rats (Table 2), but is present in primates (Table 3). ‘Knocking it out’ in rodents has no impact on blood pressure (Figure 1C). Consequently, rs6141767 itself is a byproduct of primate evolution, and a human-centered marker for the functional C3QTL3 nearby, not the QTL per se.
3.7. Closely-linked C10QTL1 and C10QTL5 of DSS rats, (22) may be a physiological human ortholog of PPM1E (protein phosphatase, Mg2+/Mn2+ dependent 1E) and a positional human ortholog of VMP1 (vacuole membrane protein 1) [2].
3.7.1. Chr 10 of DSS rats carries several QTLs by changing BP [23]. Among them (Figure 1B), distinct C10QTL1 and C10QTL5 have become relevant to humans because of the following.
In the C10QTL1-residing region of 600 kb, 3 closely-linked genes within 329 kb are Ppm1e, Rad51c (Rad51 homolog c) and Tex14 (testis expressed 14) as a genome block. Among multiple intronic GWAS SNPs, no conservation was found in the rat (Table 2). These SNPs are by products of primate evolution (Table 3). They alone or collectively do not change BP by C10QTL1 in vivo (Figure 1C), since humans and rodents achieve similar BP with or without them [10].
3.7.2. The level of C10QTL1 expression does not have a physiological impact in changing BP, since 1 copy of the normotensive C10QTL1 allele lowered BP to the same extent as 2 copies in vivo [24]. Among the 3 genes, Ppm1e possesses a missense mutation and appeared to be the principal function candidate for C10QTL1 [22]. No other structural variants [25] were found in the remaining 2 genes. It is unclear if each of PPM1E, RAD51C and TEX14 would represent a single QTL or they constitute a genome block with 1 QTL residing in it. Human data bases showed 8 probable PPM1E missense mutations (Supplementary Table 5).
3.7.3. Unlike C10QTL1/ PPM1E, Vmp1 does not carry structural mutations (Supplementary Table 6), yet an intronic SNP, rs2645466, in VMP1 was found to be associated with BP. Rs2645466 is a by-product of primate evolution (Table 3), is not conserved in the rat (Table 2) and thus does not impact on the functionality of C10QTL5/VMP1 on BP.
3.7.4. The functional candidate for C10QTL5 from the DSS rat, proline rich 11 (Prr11) bears 2 missense mutations [22]. No GWAS signal appeared near the human PRR11 (Figure 1C), which is about 400kb away from VMP1. Thus, it is likely that 2 separate QTLs may exist in the C10QTL5-residing interval.
3.7.5. In contrast to the functional correspondence of the 4 human GWAS genes to C10QTL1 and C10QTL5 (Figure 1B), there was no BP effect by knocking in the rat ortholog of a human GWAS gene, ACE, in C10S.L8. Thus, the relevance of ACE as a human GWAS gene in BP regulation needs to be tested in animal models other than DSS.
4. Discussion
Principal findings from this study are (a) in vivo studies and human GWAS have revealed shared mechanisms of BP control as a physiological framework originating from common ancestors of humans and rodents. The M3R signaling pathway is one of them. (b) Specifically, 6 distinct QTLs from inbred DSS rats have unraveled mechanistic causes in BP regulation for at least 6 new human GWAS genes. Each of them has a major physiological impact on BP, and they may collectively function in 2 pathways. (c) Previously-unsuspected components of these pathways have been implicated from the candidate genes for the QTLs. (d) The non-coding SNPs marking these 6 QTLs/human GWAS genes are offshoots of primate evolution irrelevant to BP regulation. These SNPs are human-centered and mark potential QTLs nearby, rather than being QTLs per se.
4.1.1. QTL Modularity/pathway is the physiological framework of QTLs regulating blood pressure invented in mammalian ancestors: QTL Modularity is the genetic framework in physiologically modulating BP embedded in their ancestral genomes [9]. The 6 QTLs and their corresponding human GWAS genes may function via only 2 modules in physiologically controlling BP and implicates 2 pathways of hypertension pathogeneses. One of them is the M3R signaling pathway [5,6]. This conservation of BP-regulating pathways such as the M3R signaling supports similarity in blood pressures between differing orders of mammals [10]. As a result, the fundamental framework of BP-controlling mechanisms in pathways with multiple steps must have been established in common ancestors of mammals before they started to diverge [9].
4.1.2. Humans and rodents along with most land placental and marsupial mammals diverged at various times during the past 160 million years (www.timetree.org), yet they all have similar blood pressures [10]. Environments have changed during the evolution of these mammals, and present-day humans and rodents live in very different surroundings. Convergent evolution from ancestors via different mechanisms cannot produce similar blood pressures by accident for all these orders of mammals. The only course for this to take is that key mechanisms regulating BP must have been established in common ancestors of mammals, before they evolutionarily diverged.
4.1.3. When we probe mechanisms regulating the current physiology in humans, we actually dig into the mechanistic past before humans existed. This means that, similar to those in rodents, physiological mechanisms controlling human blood pressure have already been given by ancestral QTLs in polygenic forms to virtually 100%. Nevertheless, this conservation does not exclude a later emergence of new BP-modulating mechanisms during mammalian evolution. For example, elephants and giraffes have blood pressures different from humans and rodents [10], despite sharing ancestral genomes. Humans could have evolved new mechanisms regulating BP, but all these possible developments combined and unique to humans seems to have contributed to close to 0% in the total human physiology regulating blood pressure.
4.1.4. In this context, non-coding SNPs used in GWAS are unique to humans, as a result of a coincidence in primate evolution unrelated to blood pressure control. Such a human-centered SNP fortuitously marks a physiological QTL nearby, similar to a rodent polymorphism [26] marking C17QTL1/Chrm3 next to it [5]. The paradox of a ‘common’ SNP/marker with no effect on BP identifying a ‘rare’ BP-impacting variant nearby has been previously addressed in Discussion in reference [11], and will not be reiterated here.
4.1.5. More than 10,000 human SNPs are found to be associated with over 900 genes [2]. Apparently, not all of these SNPs can be functional variants, in spite of their comparably-strong statistical significance in GWAS. Thus, genetic architectures from GWAS are no equivalent to physiological functions. Although <10% of these human-centric non-coding GWAS SNPs might potentially have functions in modulating cellular gene expressions, epigenetics and/or even be eQTLs [2], they contribute little to the primary mammalian physiology of BP controls including for humans.
4.2.1. Uncoupling between systemic BP and cell/tissue activities necessitates an in vivo physiological proof of a BP QTL: Even though a QTL is a genetic term covering a chromosome region marked by a GWAS SNP, a QTL is molecularly one gene, physiologically capable of altering BP [5,11]. Statistics is inadequate to establish this fact. Indirectly, GWAS results lead to certain studies that analyze in vitro functions of human non-coding SNPs in cells implicating in blood pressure control. However, a separation between systemic BP and cell activities casts doubt on this in vitro approach.
4.2.2. Depleting M3R diminishes vaso-relaxation that is supposed to increase BP, but contrarily decreases blood pressure [5,7]. Thus, viewing functions of these cell/tissue structures in vitro and in isolation cannot predict the actual physiological BP in vivo for a QTL. M3R is mostly produced in the brain, less in adrenals and not detectable in heart, kidneys [5,7]. We are still puzzled [6] as to how M3R promotes hypertension, and from the brain and adrenals, modulates concordant cardiac/renal functions, but discordant vaso-relaxation in vasculature. Partially inferring BP mechanisms from cells could be misleading. The systemic BP physiology in vivo is not random and detached cellular events, but integrated and offsetting interactions among various organs. No in vitro substitutes can replace the in vivo physiology in authenticating a QTL for blood pressure.
4.3.1 Genetic architectures of human GWAS designate probable locations of QTLs for functional proxies to follow in vivo. Results presented in the current manuscript are based on physiological studies in vivo from inbred rodent strains, and seem perplexing and unsettling from the perspective of the quantitative genetics principle predicating on GWAS from outbred populations [3]. However, exhibiting disparate genetic architectures between inbreds and outbreds does not imply their mechanistic differences in modulating their blood pressure physiology.
4.3.2. In outbred human populations, persons with high and low blood pressures exist, but their hypertensive and normotensive phenotypes do not become distinguishable and heritable traits until singled out and fixed as strains as by inbreeding. Inbreeding in rodents captures, but does not change, the machinery controlling BP physiological mechanisms for an individual from an outbred population. In this way, inbreedings have achieved in building hypertensive and normotensive strains the same way as Mendel was naturally given in contrasting sizes in peas from outbred populations [27]. For example, Mendel picked peas from outbred populations with explicit size differences that are heritable and contrasting. His choices were purely due to their clear and unambiguous distinctions to facilitate his phenotypings. In hindsight, if he would have analyzed continuous and ambiguous phenotypes in outbred populations, he might not have gained insights into fundamental laws of heredity.
4.3.3. By exploring inbred rat strains with heritable, contrasting and distinguishable BP features, we have unraveled mechanisms and the physiology of BP regulations from QTLs in an individual [9]. An outbred population of 100 is equivalent to 100 different inbreds in mechanisms of BP regulations. Studying all 100 in a mixture of continuous and equivocal variations is not mechanistically informative. Further in GWAS, a probable blood pressure effect from a single SNP is fractionated by total variance in phenotypic variations according to Fisher [3], not its actual physiological effect on blood pressure in mmHg.
4.4.1. Insights into mechanistic and physiological causes of blood pressure regulation from QTL modularity: The genetic modularity of QTLs [14,19] has broadened the scope of Mendelism to cover polygenic traits, and revealed causes and a mechanistic frame work of BP physiology [9]. Polygenicity of blood pressure is composed of individual Mendelian ‘monogenic’ components that are organized into modules. Mendelism is the fundamental basis for BP as a polygenic trait, and is in principle, equivalent to carbon being the basic chemical element in forming poly-carbon graphite and diamonds.
4.4.2. Recently, an ‘omnigenic’ hypothesis has been proposed to explain GWAS results on generic quantitative phenotypes [3]. It can be described as an anthropocentric (or human-centered) theory, because non-coding GWAS SNPs only exist in humans, and not in rodents, as our previous [11] and current findings have shown. It basically proposes that regulations at gene expressions at cellular level would determine the GWAS SNPs’ roles in human phenotypes including BP. This is contrary to the modularity idea and physiological proofs underlying it [9].
4.4.3. The basic distinction between the two is that modularity is predicated on the physiological causes of pathogenic mechanisms of hypertension versus statistical epidemiology that focuses on after effects of these mechanisms and physiology. This is because mechanisms and physiology determining a polygenic trait are the prime mover and starting point driving the formation of the modularity paradigm. In contrast, mechanisms and physiology are only an afterthought that the omnigenic model injects.
4.4.4. These 2 opposing hypotheses generate contrasting predictions that can be experimentally tested for their physiological relevance to BP. On the basis of functional physiology in BP regulation, modularity has been validated, but omnigenicity has not, because of the following functional results.
4.4.5. Central to the ‘omnigenic’ hypothesis is the cellular gene dose, and by inference, the level of gene expressions that would lead to a phenotype. Contrary to this prediction, several lines of experimental evidence have proven that a gene dose is irrelevant to the physiological outcome on BP control. For example, blood pressures are the same between Chrm3+/- with one functional copy of Chrm3 and Chrm3+/+ with 2 copies [5,7]. Most QTLs function in similar gene-dose independence in the physiology of BP control [24], in that one dose of a normotensive QTL allele has the same BP impact as 2 doses. There was no evidence that the level of Chrm3 expression was different in the organs tested when blood pressure changed [5].
4.4.6. The ‘miniscule’ effect from a single QTL fractionated from total variance is another prediction from the ‘omnigenic’ hypothesis. If this prediction were physiologically pertinent, removing one such QTL should have an inconsequential effect on BP. This is not the case, as introduced in Result (3.3) from gene targeting and congenic knock in experiments.
4.4.7. The modularity paradigm can, whereas the ‘omnigenic’ hypothesis cannot, explain the evolutionary conservation in pathways controlling BP rooted in common ancestors of humans and rodents. The anthropocentric non-coding GWAS SNPs that gave rise to ‘omnigenicity’ cannot affect these pathways, because they only began to appear in primates, but do not exist in rodents (Tables 2 and 3). Conversely, rodents’ non-coding Chrm3 SNPs are not conserved in humans. Inactivating the M3R signaling pathway did not touch any of them, yet BP changed [5]. Thus, the functional M3R signaling coexists in humans and rodents, and should be shared in determining the hypertension pathogenesis, to which the non-coding rodent Chrm3 SNPs and the human-centered GWAS SNPs are irrelevant.
4.4.8. Certain QTLs starts to function at embryogenesis [5], before the onset of adult BP physiology. Modularity can [9], whereas the ‘omnigenicity’ cannot, explain that a pathway involved in BP control can temporally begin at embryogenesis and continue into adulthood [5].
4.4.9. In conclusion, modularity is supported by reproducible lines of physiological evidence as a signaling pathway, and is mechanistic in our understandings of causes for BP physiology [9]. The intuitive ‘omnigenic’ hypothesis [3] has little functional support for a physiological role on BP, despite explaining epidemiological exhibitions from human GWAS. It’s analogous to Newtonian mechanics describing effects of gravity as a force pulling on objects. When it comes to the cause of gravity, we have to switch our mindset to Einstein’s theory of a curvature in space time for explanation (https://www.britannica.com/science/general-relativity).
4.5.1. Pathogenic pathways of hypertension inferred from the molecular bases of QTLs: Since the 3 following QTLs (Figure 1, Table 1) have not been molecularly identified like C17QTL1/Chrm3, their roles in BP physiology are tentatively inferred from the functional candidate genes representing them.
4.5.2. In epistatic module 2/M3R signaling pathway [5,7], RREB1 is ras responsive element binding protein 1, a transcription factor involved various molecular processes and implicated in certain diseases [28]. Its probable mechanistic step in the M3R signaling pathway remains to be investigated.
4.5.3. In epistatic module 1/pathway 1, 2 new components are suggested, i.e. DNMT3B and PPM1E (Table 1). Their functions begin during embryogenesis [29]. DNMT3B encodes the DNA methyltransferase 3B whose defects cause human Immunodeficiency, Centromeric instability and Facial anomalies (ICF) (30). The DNMT3B enzyme methylates distinct CpG islands de novo in embryonic stem cells [29]. 2 Dnmt3b missense mutations, at A609T and D823G amino acids, seem to hypo-methylate repetitive DNA sequences and resemble the human ICF syndrome in certain phenotypes [31].
4.5.4. PPM1E is a protein phosphatase, Mg2+/Mn2+ dependent 1E [32] principally expressed in the brain [33], and belongs to a family of serine/threonine-protein phosphatases. Very little is known of its function in vivo, since no knock out exists presumably due to its requirement during development.
4.6.1. Caveats and limitations are: First, although human GWAS genes with missense mutations do not genetically prove by themselves to be the QTLs in questions, they provide entry points towards probable steps in 2 pathways contributed by each QTL. Molecular designs with viable gene-targeting of their codons [5] can test their functions on blood pressure in rodents.
4.6.2. Second, structural mutations are not necessarily the only molecular bases that can affect a pathway in question. C10QTL5/VMP1 has no missense mutation in DSS rats. Despite of it, it can still be a BP QTL, because certain steps in epistatic module 1/pathway 1 may depend on a multi-component complex. One component can affect the function of the entire complex by altering its stoichiometry of the composition. As a result, the pathway as a whole is affected.
4.6.3. Finally, unlike the small segment harboring C17QTL1/Chrm3, the regions containing the remaining 5 QTLs are quite large (Figure 1). The number of QTLs in each of the 5 regions is under reported. At least 3 QTLs exist in the C17QTL2-residing interval. 2 QTLs at minimum lie in the segment bearing C3QTL3 [34,35]. A minimum of 3 QTLs may be present in the C3QTL1-lodging region, because human GWAS genes exist on 3 separate CHROMOSOMES (Supplementary Table 1).
5. Conclusions
Studies of human epidemiology and animal models in polygenic hypertension have been accidentally divided into 2 practices and doctrines that conveniently and separately govern each. However, this artificial partition is not based on the physiology and mechanisms, and cannot hide an inconvenient truth that both mammalian orders share fundamental mechanisms in BP controls at least 90 million years in the making. The reproducible experimental evidence has reinforced the verdict that shared BP QTLs are causes of these mechanisms. QTL modularity mechanistically joins humans and rodents, suggests that multiple QTLs may function in a common pathway, and each is involved in a different step in the pathogenic pathway towards polygenic hypertension. This emerging paradigm encompasses not only humans, but also most other land mammals, and is a departure from the human-centric precept [3] which is reminiscent of geocentrism distorting heliocentricy of our solar system in cosmology.
Funding
The work was supported by a grant from Heart and Stroke Foundation of Canada to AYD.
Acknowledgements
We appreciate the support of CRCHUM cardiovascular phenotyping core facility.
Conflict of interest
None.
References
- Go AS, Mozaffarian D, Roger VL, et al. Heart Disease and Stroke Statistics 2013 Update: A Report From the American Heart Association. Circulation 127 (2013): e6-e245.
- Evangelou E, Warren HR, Mosen-Ansorena D, et al. Genetic analysis of over 1 million people identifies 535 new loci associated with blood pressure traits. Nature Genetics 50 (2018): 1412-25.
- Boyle EA, Li YI, Pritchard JK. An Expanded View of Complex Traits: From Polygenic to Omnigenic. Cell 169 (2017): 1177-86.
- Deng AY. Positional Cloning of Quantitative Trait Loci for Blood Pressure: How Close Are We?: A Critical Perspective. Hypertension 49 (2007): 740-7.
- Deng AY, deBlois D, Laporte SA, et al. Novel Pathogenesis of Hypertension and Diastolic Dysfunction Caused by M3R (Muscarinic Cholinergic 3 Receptor) Signaling. Hypertension 72 (2018): 755-64.
- Cowley AW. Chrm3 Gene and M3 Muscarinic Receptors Contribute to Salt-Sensitive Hypertension, But Now a Physiological Puzzle. Hypertension 72 (2018): 588-91.
- Deng AY, Huot-Marchard J-É, deBlois D, Thorin E, Chauvet C, Menard A. Functional Dosage of Muscarinic Cholinergic Receptor 3 Signalling, Not the Gene Dose, Determines Its Hypertension Pathogenesis. Canadian Journal of Cardiology 35 (2019): 661-70.
- Alves-Lopes R, Neves KB, Touyz RM. Muscarinic Receptor Type-3 in Hypertension and Cholinergic-Adrenergic Crosstalk: Genetic Insights and Potential for New Antihypertensive Targets. Canadian Journal of Cardiology 35 (2019): 555-7.
- Deng AY. Modularity/non-cumulativity of quantitative trait loci on blood pressure. J Hum Hypertens 34 (2020): 432-9.
- White CR, Seymour RS. The role of gravity in the evolution of mammalian blood pressure Evolution 68 (2014): 901-8.
- Deng AY, Ménard A. Biological convergence of three human and animal model quantitative trait loci for blood pressure. Journal of Hypertension 38 (2020): 322–31
- Deng AY, Ménard A. Functional Captures of Multiple Human Quantitative Trait Loci Regulating Blood Pressure with the Use of Orthologs in Genetically Defined Rat Models. Canadian Journal of Cardiology 36 (2020): 756-63.
- Consortium W. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447 (2007): 661-78.
- Chauvet C, Crespo K, Menard A, Roy J, Deng AY. Modularization and epistatic hierarchy determine homeostatic actions of multiple blood pressure quantitative trait loci. Human Molecular Genetics 22 (2013): 4451-9.
- Deng AY. Genetic basis of polygenic hypertension. Human Molecular Genetics. 2007;16(R2):R195-R202.
- Charron S, Lambert R, Eliopoulos V, Duong C, Menard A, Roy J, et al. A loss of genome buffering capacity of Dahl salt-sensitive model to modulate blood pressure as a cause of hypertension. Hum Mol Genet 14 (2005): 3877-84.
- Crespo K, Menard A, Deng AY. Hypertension Suppression, Not a Cumulative Thrust of Quantitative Trait Loci, Predisposes Blood Pressure Homeostasis to Normotension. Circulation: Cardiovascular Genetics 8 (2015): 610-7.
- Harrap SB, Morris BJ. Blood Pressure Genetics Just Don't Add Up. Circ Cardiovasc Genet 8 (2015): 541-3.
- Deng AY. Genetic mechanisms of polygenic hypertension: fundamental insights from experimental models. Journal of Hypertension 33 (2015): 669-80.
- Garrett MR, Dene H, Walder R, et al. Genomic scan and congenic strains for blood pressure quantitative trait loci using Dahl salt-sensitive rats. Genome Research 8 (1998): 711-23.
- The Genomes Project C, Auton A, Abecasis GR, et al. A global reference for human genetic variation. Nature 526 (2015): 68-74.
- Chauvet C, Menard A, Xiao C, et al. Novel genes as primary triggers for polygenic hypertension. J Hypertens 30 (2012): 81-6.
- Chauvet C, Charron S, Menard A, Xiao C, Roy J, Deng AY. Submegabase resolution of epistatically interacting quantitative trait loci for blood pressure applicable for essential hypertension. J Hypertens 26 (2008): 893-901.
- Duong C, Charron S, Deng Y, et al. Individual QTLs controlling quantitative variation in blood pressure inherited in a Mendelian mode. Heredity 98 (2007): 165-71.
- Sudmant PH, Rausch T, Gardner EJ, et al. An integrated map of structural variation in 2,504 human genomes. Nature 526 (2015): 75-81.
- Deng AY, Dene H, Pravenec M, Rapp JP. Genetic mapping of two new blood pressure quantitative trait loci in the rat by genotyping endothelin system genes. J Clin Invest 93 (1994): 2701-9.
- Birchler JamesA. Mendel, Mechanism, Models, Marketing, and More. Cell 163 (2015): 9-11.
- Deng Y-N, Xia Z, Zhang P, Ejaz S, Liang S. Transcription Factor RREB1: from Target Genes towards Biological Functions. International Journal of Biological Sciences 16 (2020): 1463-73.
- Okano M, Bell DW, Haber DA, Li E. DNA Methyltransferases Dnmt3a and Dnmt3b Are Essential for De Novo Methylation and Mammalian Development. Cell 99 (1999): 247-57.
- Xu G-L, Bestor TH, Bourc'his D, et al. Chromosome instability and immunodeficiency syndrome caused by mutations in a DNA methyltransferase gene. Nature 402 (1999): 187-91.
- Ueda Y, Okano M, Williams C, Chen T, Georgopoulos K, Li E. Roles for Dnmt3b in mammalian development: a mouse model for the ICF syndrome. Development 133 (2006): 1183-92.
- Voss M, Paterson J, Kelsall IR, et al. Ppm1E is an in cellulo AMP-activated protein kinase phosphatase. Cellular Signalling 23 (2011): 114-24.
- Magdaleno S, Jensen P, Brumwell CL, et al. BGEM: an in situ hybridization database of gene expression in the embryonic and adult mouse nervous system. PLoS Biol 4 (2006): e86-e.
- Lee SJ, Liu J, Westcott AM, et al. Substitution mapping in dahl rats identifies two distinct blood pressure quantitative trait loci within 1.12- and 1.25-mb intervals on chromosome 3. Genetics 174 (2006): 2203-13.
- Koh-Tan HHC, Dashti M, Wang T, et al. Dissecting the genetic components of a quantitative trait locus for blood pressure and renal pathology on rat chromosome 3. Journal of hypertension 35 (2017): 319-29.
- Chauvet C, Menard A, Deng AY. Two candidate genes for two quantitative trait loci epistatically attenuate hypertension in a novel pathway. J Hypertens 33 (2015): 1791-801.
- Atanur S-á, Diaz A-á, Maratou K, et al. Genome Sequencing Reveals Loci under Artificial Selection that Underlie Disease Phenotypes in the Laboratory Rat. Cell 154 (2013): 691-703.
This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license 4.0
Supplementary Table 1: Rat QTLs and genes residing in the intervals harboring them that correspond to human GWAS genes trapped by congenic knock in genetics.
Footnote for table: Fig. 1 of the text defines the chromosome regions containing QTLs and the gene residing in the intervals containing these QTLs. Human GWAS genes are all from (Nat Genet 2018;50:1412), except for CHRM3 marked by rs2820037, which was taken from (Nature 2007;447:661). C17QTL1/Chrm3 is adapted from (Hypertension 2018; 72:755). Only the genes underlined were analyzed further in the current work, because they are functional candidate genes for the human QTLs. CHRM3, muscarinic cholinergic receptor 3; DNMT3B, DNA methyltransferase 3 beta; PPM1E, protein Phosphatase, Mg2+/Mn2+ Dependent 1E; RREB1, ras responsive element binding protein 1; VMP1, Vacuole membrane protein 1. CHR, Chromosome.
Supplementary Table 2. Amino acid alignment for muscarinic cholinergic receptor 3 (M3R) among Tasmania devils (marsupial mammals), humans and rats (both placental mammals).
Footnote for table: * denotes an overall amino acid identity (92.3%) between humans and rats, (90.8%) between humans and Tasmania Devils and (90.3%) between rats and Tasmania Devils. Underlined codons indicate the domain involved in M3R signaling. The single missense mutation in DSS rat is shaded at M and alters the M3R signaling, and is associated with BP changes. Probable human M3R missense mutations are marked and curated from https://www.ncbi.nlm.nih.gov/snp/?term=chrm3+missense. Only those human mutations with minor alleles that were observed at least 2 times in the tested populations are given.
Supplementary Information:
Supplementary Table 3: Mutation survey of genes in rat QTL-residing intervals that capture human GWAS genes by congenic knock in genetics
Footnote to Table: Gene locations are indicated on the map in Fig. 1. The position of a mutation enumerates from the ATG start codon of that gene. The amino acid position begins from the first methionine. Confirmed mutations are indicated by bold letters. DNMT3B, DNA methyltransferase 3 beta; PPM1E, protein phosphatase, Mg2+/Mn2+ dependent 1E; RREB1, ras responsive element binding protein 1; No Copy Number Variation (CNV) had been found for those genes from total genome sequencing of DSS and Lewis rats based on our current work and those of the rat genome database (RGD).
Supplementary Table 4: Amino acid alignment and missense mutations in codons DNA for methyltransferase 3 beta (DNMT3B) in humans and rats
Footnote for table: * indicates amino acid identity (92%) between human and the rat. Probable human missense mutations (The Genomes Project et al. 2015) are shaded, which were curated from https://www.ncbi.nlm.nih.gov/snp/ and as of April 23, 2020. Only those missense mutations with minor alleles that were observed at least 2 times (marked in red) in the tested populations are included with the validation status by 1000Genomes. Amino acids in blue indicate that the minor allele has been observed more than 10 times. The rat missense mutation is shaded in green. DSS, Dahl salt-sensitive rats.
Supplementary Table 5: Amino acid alignment and missense mutations in codons for protein phosphatase, Mg2+/Mn2+ dependent 1E (PPM1E) in humans and rats
Footnote for table: * indicates amino acid identity (92%) between human and the rat. Probable human missense mutations (The Genomes Project et al., 2015) are shaded, which were curated from https://www.ncbi.nlm.nih.gov/snp/?term=ppm1e+missense and as of Nov. 12, 2018. Only those missense mutations with minor alleles that were observed at least 2 times (marked in red) in the tested populations are included with the validation status by 1000Genomes. Amino acids in blue indicate that the minor allele has been observed more than 10 times. The rat missense mutation is shaded in green. DSS, Dahl salt-sensitive rats.
Supplementary Table 6: Amino acid alignment and missense mutations in codons DNA for vacuole membrane protein 1 (VMP1) in humans and rats
Footnote for table: * indicates amino acid identity (96%) between human and the rat. Probable human missense mutations (The Genomes Project et al. 2015) are shaded, which were curated from https://www.ncbi.nlm.nih.gov/snp/ and as of April 23, 2020. Only those missense mutations with minor alleles that were observed at least 2 times (marked in red) in the tested populations are included with the validation status by 1000Genomes. Amino acids in blue indicate that the minor allele has been observed more than 10 times. DSS, Dahl salt-sensitive rats.